External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G53170 97 / 3e-25
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G48730 71 / 5e-16
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G35130 52 / 2e-09
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT2G41720 51 / 6e-09
EMB2654
EMBRYO DEFECTIVE 2654, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G18900 49 / 2e-08
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2), Pentatricopeptide repeat (PPR) superfamily protein (.3)
AT1G74750 48 / 7e-08
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62720 48 / 9e-08
AtNG1
novel gene 1, Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT4G11690 46 / 3e-07
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT3G06430 45 / 5e-07
AtPPR2, EMB2750
pentatricopeptide repeat 2, embryo defective 2750, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63080 44 / 1e-06
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024893
155 / 7e-47
AT3G53170 543 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10022787
69 / 4e-15
AT5G48730 687 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10011849
69 / 4e-15
AT5G48730 681 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10035818
56 / 2e-10
AT4G30825 1049 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10036600
55 / 3e-10
AT4G30825 428 / 3e-144
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10042068
52 / 4e-09
AT2G41720 1057 / 0.0
EMBRYO DEFECTIVE 2654, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10018075
52 / 4e-09
AT2G41720 1020 / 0.0
EMBRYO DEFECTIVE 2654, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10025473
51 / 7e-09
AT2G18940 970 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10006962
49 / 3e-08
AT2G18940 976 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G058300
112 / 1e-30
AT3G53170 618 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G243600
79 / 9e-19
AT5G48730 691 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G153900
56 / 2e-10
AT3G06430 717 / 0.0
pentatricopeptide repeat 2, embryo defective 2750, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.012G123600
52 / 2e-09
AT2G35130 852 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.008G098700
51 / 8e-09
AT3G06430 692 / 0.0
pentatricopeptide repeat 2, embryo defective 2750, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G030200
50 / 9e-09
AT1G19720 985 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.006G166200
48 / 7e-08
AT2G18940 1082 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G084000
47 / 1e-07
AT3G22670 489 / 3e-168
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.006G141800
47 / 2e-07
AT2G06000 588 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1), Pentatricopeptide repeat (PPR) superfamily protein (.2)
Potri.019G043101
47 / 2e-07
AT5G14770 796 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10024144 pacid=23141598 polypeptide=Lus10024144 locus=Lus10024144.g ID=Lus10024144.BGIv1.0 annot-version=v1.0
ATGAGGCATGTGGAGAATTCGGATGTGGTTCTGGATACTCCTTTCTTTAACTGTGTCATCAGTGCTTATGGTCGTGCTGGAGAAGTGGAAAACATGGAAG
GGTTGTTTTCGAGCATGGCAGAGAAGAATTGCAGGCCTGATTATGTCGCGTACGCTACTATGATCCAGGCTTACAATGCGAGAGGAATGACGGACCCTGC
AATAGATTTGGAAAAGAAGATGCTTGCAGCACAAAAAAGATCAGGTATCTGA
AA sequence
>Lus10024144 pacid=23141598 polypeptide=Lus10024144 locus=Lus10024144.g ID=Lus10024144.BGIv1.0 annot-version=v1.0
MRHVENSDVVLDTPFFNCVISAYGRAGEVENMEGLFSSMAEKNCRPDYVAYATMIQAYNARGMTDPAIDLEKKMLAAQKRSGI
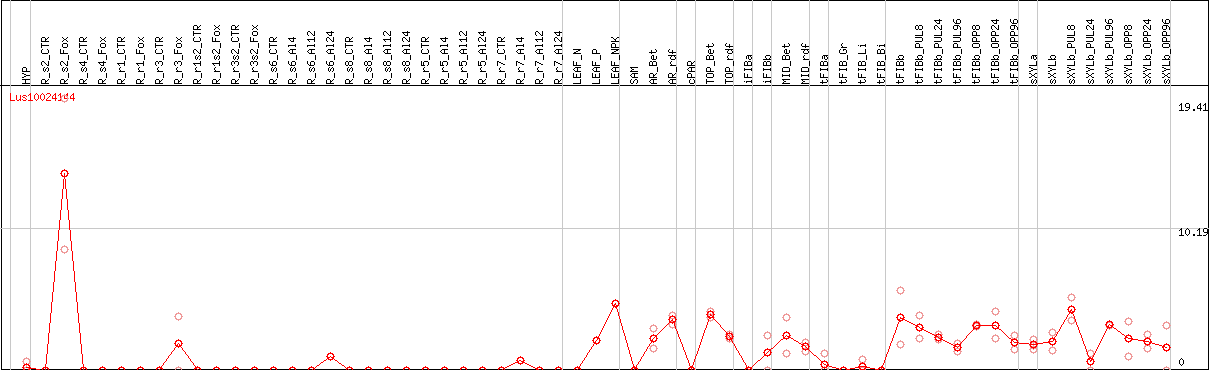
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024144 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.