External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G47800 155 / 3e-46
Galactose mutarotase-like superfamily protein (.1)
AT3G17940 114 / 9e-31
Galactose mutarotase-like superfamily protein (.1)
AT5G15140 111 / 7e-29
Galactose mutarotase-like superfamily protein (.1)
AT3G01260 72 / 8e-15
Galactose mutarotase-like superfamily protein (.1)
AT3G47780 56 / 2e-09
ABCA7, ATATH6
A. THALIANA ABC2 HOMOLOG 6, ATP-binding cassette A7, ABC2 homolog 6 (.1)
AT3G47760 52 / 3e-08
ATATH4, ABCA5
A. THALIANA ABC2 HOMOLOG 4, ATP-binding cassette A5, ABC2 homolog 4 (.1)
AT3G47790 52 / 3e-08
ABCA8, ATATH7
ATP-binding cassette A8, ABC2 homolog 7 (.1)
AT5G61740 52 / 5e-08
ABCA10, ATATH14
ARABIDOPSIS THALIANA ABC2 HOMOLOG 14, ATP-binding cassette A10, ABC2 homolog 14 (.1)
AT5G61700 51 / 8e-08
ABCA12, ATATH16
ARABIDOPSIS THALIANA ABC2 HOMOLOG 16, ATP-binding cassette A12, ABC2 homolog 16 (.1)
AT3G47770 51 / 1e-07
ATATH5, ABCA6
A. THALIANA ABC2 HOMOLOG 5, ATP-binding cassette A6, ABC2 homolog 5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023611
237 / 2e-78
AT3G47800 462 / 5e-164
Galactose mutarotase-like superfamily protein (.1)
Lus10005251
131 / 2e-37
AT5G15140 370 / 2e-126
Galactose mutarotase-like superfamily protein (.1)
Lus10015365
130 / 2e-36
AT5G15140 403 / 7e-138
Galactose mutarotase-like superfamily protein (.1)
Lus10030673
131 / 3e-36
AT5G15140 358 / 1e-118
Galactose mutarotase-like superfamily protein (.1)
Lus10007251
129 / 3e-36
AT5G15140 404 / 7e-139
Galactose mutarotase-like superfamily protein (.1)
Lus10010976
115 / 3e-31
AT3G17940 497 / 4e-178
Galactose mutarotase-like superfamily protein (.1)
Lus10026181
97 / 3e-24
AT3G17940 500 / 3e-179
Galactose mutarotase-like superfamily protein (.1)
Lus10000452
87 / 8e-21
AT3G17940 436 / 1e-154
Galactose mutarotase-like superfamily protein (.1)
Lus10023612
0 / 1
AT3G47780 1202 / 0.0
A. THALIANA ABC2 HOMOLOG 6, ATP-binding cassette A7, ABC2 homolog 6 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0103
Gal_mutarotase
PF01263
Aldose_epim
Aldose 1-epimerase
Representative CDS sequence
>Lus10024242 pacid=23175783 polypeptide=Lus10024242 locus=Lus10024242.g ID=Lus10024242.BGIv1.0 annot-version=v1.0
ATGTCATTTTCTCTGACTTTTGCATCGGGAAAAGGTAGAGCAATTGGTACTAGAACCAAATACAAGTCATGCGATAGTCTGTACAACCTCATAAAGGTGT
ATCCAGGGAGGGCTGGGAATCAAGCCAATTTGGCGGTGAAAGGATTATTGCTTGCGCTTCCTCGAGGGGAGTGTTTTGGGAACCTCGGCGGACACAACAG
TGGAACCATCCTCAACCACACCATCCATCTCTTCTCCCACCACATCACTCCTGTCAACGACCACCTCATCCCTACCGGATCTTTGACTTCTGTAGAGGGC
ACCCCTTACGACTTCCTTGAGCCTCTACAAATCGGTAGCAGGATCGACCGTGATTTCCCCAAAGGGTACGACATTAACTACGTGCTGGATGACCTTTCTC
CAGGCCACCTAAAGAAAGCTGCTGTTGTTAAGGATCATGTCTCAGGGAGGAAACTTGAGCTGTGGACTAATCAGCCTGGAGTCCAGTTTTATACTAGCAA
TATGTTGAACGATGTCAAGGGGAAAGATGGGTTTACTTATTAG
AA sequence
>Lus10024242 pacid=23175783 polypeptide=Lus10024242 locus=Lus10024242.g ID=Lus10024242.BGIv1.0 annot-version=v1.0
MSFSLTFASGKGRAIGTRTKYKSCDSLYNLIKVYPGRAGNQANLAVKGLLLALPRGECFGNLGGHNSGTILNHTIHLFSHHITPVNDHLIPTGSLTSVEG
TPYDFLEPLQIGSRIDRDFPKGYDINYVLDDLSPGHLKKAAVVKDHVSGRKLELWTNQPGVQFYTSNMLNDVKGKDGFTY
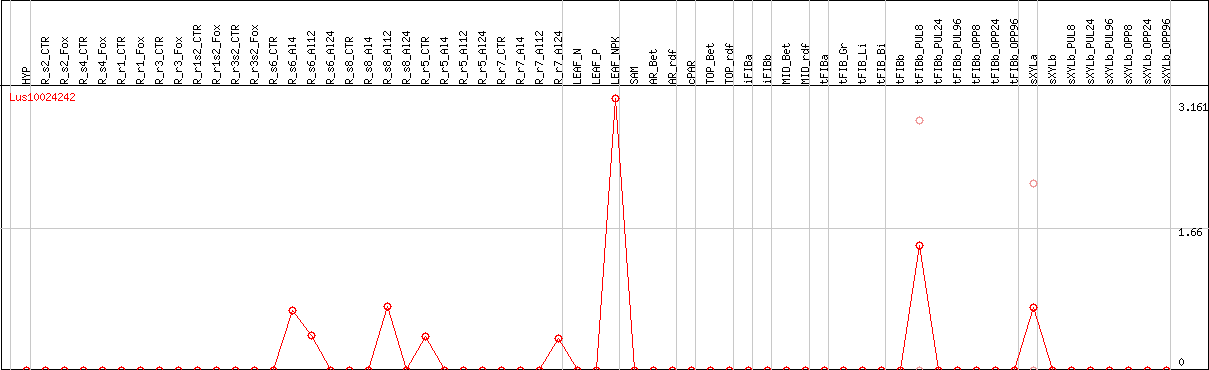
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024242 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.