External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G24503 208 / 2e-65
ALDH1A, REF1, ALDH2C4
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
AT3G48000 147 / 8e-42
ALDH2A, ALDH2B4
aldehyde dehydrogenase 2A, aldehyde dehydrogenase 2B4 (.1)
AT1G23800 142 / 3e-40
ALDH2B7
aldehyde dehydrogenase 2B7 (.1)
AT1G74920 105 / 1e-26
ALDH10A8
aldehyde dehydrogenase 10A8 (.1.2)
AT3G48170 103 / 7e-26
ALDH10A9
aldehyde dehydrogenase 10A9 (.1)
AT1G79440 96 / 3e-23
ENF1, SSADH1, ALDH5F1
SUCCINIC SEMIALDEHYDE DEHYDROGENASE 1, SUCCINIC SEMIALDEHYDE DEHYDROGENASE, ENLARGED FIL EXPRESSION DOMAIN 1, aldehyde dehydrogenase 5F1 (.1)
AT3G66658 63 / 1e-11
ALDH22A1
aldehyde dehydrogenase 22A1 (.1.2)
AT2G24270 57 / 1e-09
ALDH11A3
aldehyde dehydrogenase 11A3 (.1.2.3.4)
AT1G54100 55 / 4e-09
ALDH7B4
aldehyde dehydrogenase 7B4 (.1.2)
AT2G14170 51 / 1e-07
ALDH6B2
aldehyde dehydrogenase 6B2 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023626
252 / 6e-82
AT3G24503 753 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10019387
224 / 6e-70
AT3G24503 548 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10023625
207 / 5e-65
AT3G24503 749 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10024259
204 / 1e-63
AT3G24503 745 / 0.0
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10030143
152 / 4e-45
AT1G79580 279 / 1e-91
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
Lus10043245
148 / 5e-43
AT3G24503 487 / 2e-170
REDUCED EPIDERMAL FLUORESCENCE1, aldehyde dehydrogenase 1A, aldehyde dehydrogenase 2C4 (.1)
Lus10029166
145 / 4e-41
AT1G23800 886 / 0.0
aldehyde dehydrogenase 2B7 (.1)
Lus10012999
141 / 1e-39
AT1G23800 889 / 0.0
aldehyde dehydrogenase 2B7 (.1)
Lus10004238
106 / 7e-28
AT1G74920 533 / 0.0
aldehyde dehydrogenase 10A8 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0099
ALDH-like
PF00171
Aldedh
Aldehyde dehydrogenase family
Representative CDS sequence
>Lus10024260 pacid=23175728 polypeptide=Lus10024260 locus=Lus10024260.g ID=Lus10024260.BGIv1.0 annot-version=v1.0
ATGCAGGCAACAGCAACCAGTAATCTGAAACAAGTTTCACTTGAATTGGGTGGAAAATCTCCTCTCTTAATCTTCAACGATGCCGACATCGATGCAGCTG
CTGATCTCGCTCTCCTAGGTATCCTATTCAACAAGGGTGAAGTTTGTGTGGCAAGTTCGAGAGTCTTTGTACAAGAAGGGATATATGGCGAGTTTGTGAC
CAAATTGGTTGAAAAAGTCAAAGATTGGGTGGTTGGGGATCCTTCCAACCTCAATTCTCGTCTCGGTCCACAGGTTGATAAGAAGCAATATGAGAAAATA
ATGTCCTACATTGAGCACGGGAAAAGACAAGGAGCCACCTTGGAGGATATGATGGTAGCCCAGGACGAAATCTTTGGGCCAGTAATGGCACTAATGAAGT
TCAAAACTGTGGAGGATGCGATAAAATCGGCCAACAACACGAGGTATGGATTAGCAGCAGGGATAGTGACCAGGAACATGGACATATCAAACACTGGTGG
CAAGATCTATAAGAGCAGGAGTGATATGGATAAACTGTTACTTTGCATGTGA
AA sequence
>Lus10024260 pacid=23175728 polypeptide=Lus10024260 locus=Lus10024260.g ID=Lus10024260.BGIv1.0 annot-version=v1.0
MQATATSNLKQVSLELGGKSPLLIFNDADIDAAADLALLGILFNKGEVCVASSRVFVQEGIYGEFVTKLVEKVKDWVVGDPSNLNSRLGPQVDKKQYEKI
MSYIEHGKRQGATLEDMMVAQDEIFGPVMALMKFKTVEDAIKSANNTRYGLAAGIVTRNMDISNTGGKIYKSRSDMDKLLLCM
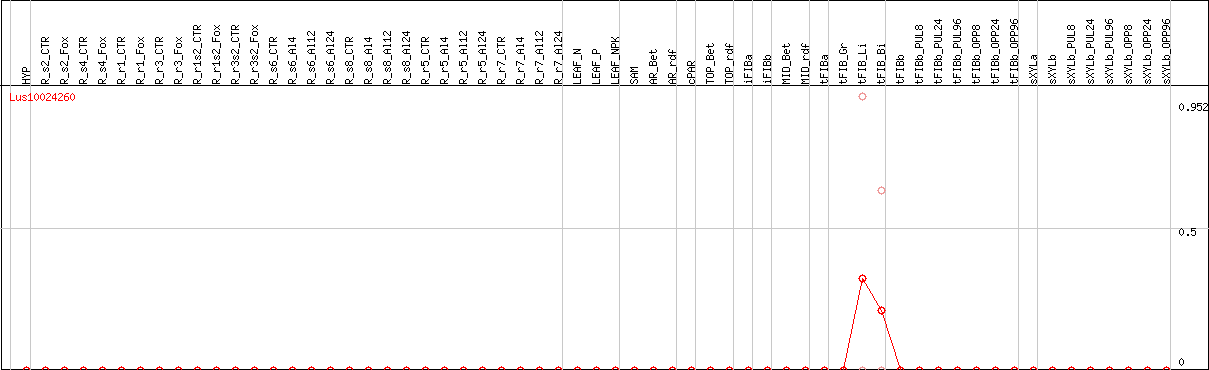
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024260 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.