External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G19630 592 / 0
CYP722A1
"cytochrome P450, family 722, subfamily A, polypeptide 1", cytochrome P450, family 722, subfamily A, polypeptide 1 (.1)
AT4G19230 243 / 4e-75
CYP707A1
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
AT5G45340 242 / 8e-75
CYP707A3
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
AT3G19270 236 / 3e-72
CYP707A4
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
AT3G13730 234 / 3e-71
CYP90D1
"cytochrome P450, family 90, subfamily D, polypeptide 1", cytochrome P450, family 90, subfamily D, polypeptide 1 (.1)
AT2G29090 226 / 1e-68
CYP707A2
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
AT5G05690 223 / 2e-67
CBB3, DWF3, CYP90A1, CYP90A, CPD
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
AT4G36380 219 / 2e-65
ROT3
ROTUNDIFOLIA 3, Cytochrome P450 superfamily protein (.1)
AT1G12740 210 / 2e-62
CYP87A2
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
AT3G30180 207 / 1e-61
CYP85A2, BR6OX2
brassinosteroid-6-oxidase 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018898
309 / 3e-100
AT5G45340 301 / 3e-97
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10028594
305 / 1e-98
AT5G45340 299 / 1e-96
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10035685
252 / 2e-78
AT4G19230 714 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
Lus10033308
249 / 2e-77
AT5G45340 732 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10014285
240 / 6e-75
AT1G19630 249 / 2e-78
"cytochrome P450, family 722, subfamily A, polypeptide 1", cytochrome P450, family 722, subfamily A, polypeptide 1 (.1)
Lus10034768
241 / 5e-74
AT5G45340 731 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10042652
233 / 6e-71
AT3G19270 647 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Lus10033105
228 / 4e-69
AT1G05160 305 / 2e-98
ENT-KAURENOIC ACID OXYDASE 1, "cytochrome P450, family 88, subfamily A, polypeptide 3", cytochrome P450, family 88, subfamily A, polypeptide 3 (.1)
Lus10024495
223 / 2e-67
AT1G12740 751 / 0.0
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G033000
610 / 0
AT1G19630 607 / 0.0
"cytochrome P450, family 722, subfamily A, polypeptide 1", cytochrome P450, family 722, subfamily A, polypeptide 1 (.1)
Potri.002G069600
306 / 2e-99
AT5G45340 296 / 1e-95
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Potri.001G242600
237 / 2e-72
AT2G29090 652 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
Potri.004G235400
235 / 4e-72
AT4G19230 778 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
Potri.014G029100
229 / 1e-69
AT3G19270 665 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.009G101700
225 / 4e-68
AT3G19270 711 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.004G140900
224 / 6e-68
AT3G19270 731 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.009G033900
222 / 6e-67
AT2G29090 677 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
Potri.002G126100
221 / 2e-66
AT3G19270 666 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.008G067500
219 / 5e-66
AT5G05690 710 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00067
p450
Cytochrome P450
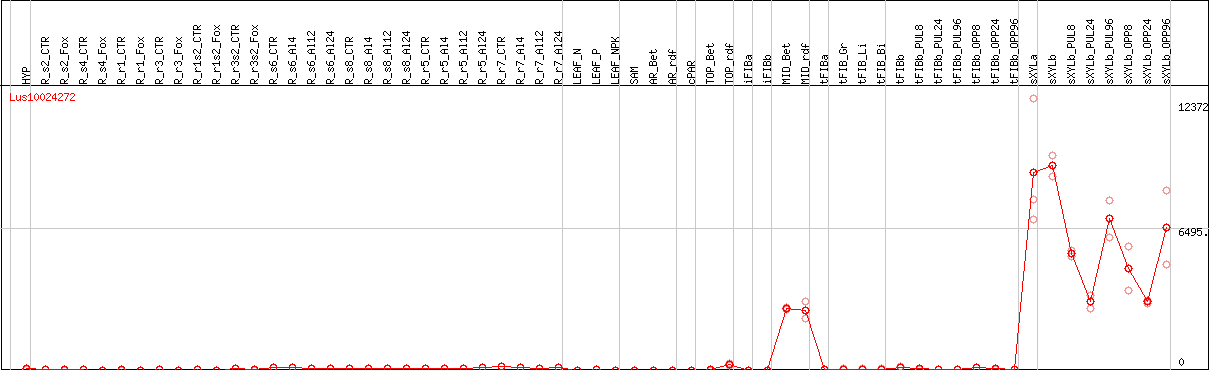
Representative CDS sequence
>Lus10024272 pacid=23175043 polypeptide=Lus10024272 locus=Lus10024272.g ID=Lus10024272.BGIv1.0 annot-version=v1.0
ATGGAAACTACAAACCTCTGTTTTGCTATCTGCGCAATACTTGTTCTTATACCACACCTACTATGGCGAAAACTCAAGATTCGACACCATAATCCCCCGC
CAGGCAGTCTTGGGTTTCCCCTCATCGGAGAGACTATACAGTTCATGGCCGCAATCAACAGCAATCGAGGATTCTACGACTTCGTCCGAGTTCGCAGCCT
AAGGTACGGGAGTTGCTTCAAGACTAACATATTTGGTTCGACTCACGTTTTTGTATCGACTACTGAATCGGCCAGGACGATCCTGAATAACGAGTCGGGT
AAATTCAGTAAGCGGTACATATGTTCGATAGGTGAGCTAGTGGGAGAGAGTAGCGTACTGTGCGCACCTTCTCAACAACACCACAAGCTGATACGGAGCC
GCCTCATGAGTTTGGTCTCCCCAACTTCATTGGGGACTTTTATCACACAGTTTGATCAGCAGATTGTCCAAAGCTTGGGGAGTTGGGAGAATGATACCAC
CGTAATTCTACTGGACCAGGCCGTCCAGATAACATTGGAGGCTATGTGCAAACTGTTGATGGGGATAGAAGAAGGAGAAGAGATAAAGAGTGTGTATGAG
GATGTATCAGACATCTTTAGAGCAATGCTGGCATTTCCTTTGAGGCTTCCTTGGACTAGATTTCATAAAGGCCTTAAGGCCCGTGAAAGAGTTATGAAGA
GACTAGATAGAATGATGTCTGAAAGGAGGGAAATCCCAGCAGATGATGAACAAGCTGACTTTCTTCAGCAGCTACTTGGAAGTTCCAACTTAACAGATGG
GGAAATTAAGGATAATATACTGACCATGATCATTGCAGGACAAGATACCACTGCAAGTGCCATGACTTGGATGGTCAAGTACCTCGCTGAAAATTCAGAT
GTTGCCCATTTATTAAGGGCTGAACAACTTCATATTTTGAAACGAAGATCAGGGAGGGAGTTCTTGGTTGCGGAAGATTTGAATGATATGCCATATGCTA
CCAAGGTTATCAAGGAGTCGCTAAGGATGGCTTCTATTGTACCATGGTTCCCTCGACTAGCACGCCAAGATTGTCAGATTGAAGGATACAAGATAAAGAA
GGGATGGAACATAAACGTTGACGCAAGATCAATTCACTTGGACCCCAAAGTATACAAGGAACCAACGAAATTCAATCCATCCAGATTTGATGATGAGGGG
ATTAGCAGTTGTTTCTTGGGATTTGGAATGGGAGGGAGGACTTGCTTGGGAATCAACTTGGCTAAATCCATGTTACTTGTTTTCCTCCATCGTCTCCTCA
CTTCTTACAAGTGGAAACTGGTGGATTCAGATTCAAGCATTGAGAAGTGGGCAATGTTCTCAAGGCTGAAAAACGGCTGCCCCATCCATTTGACGAGGAT
TACTACACCTTAA
AA sequence
>Lus10024272 pacid=23175043 polypeptide=Lus10024272 locus=Lus10024272.g ID=Lus10024272.BGIv1.0 annot-version=v1.0
METTNLCFAICAILVLIPHLLWRKLKIRHHNPPPGSLGFPLIGETIQFMAAINSNRGFYDFVRVRSLRYGSCFKTNIFGSTHVFVSTTESARTILNNESG
KFSKRYICSIGELVGESSVLCAPSQQHHKLIRSRLMSLVSPTSLGTFITQFDQQIVQSLGSWENDTTVILLDQAVQITLEAMCKLLMGIEEGEEIKSVYE
DVSDIFRAMLAFPLRLPWTRFHKGLKARERVMKRLDRMMSERREIPADDEQADFLQQLLGSSNLTDGEIKDNILTMIIAGQDTTASAMTWMVKYLAENSD
VAHLLRAEQLHILKRRSGREFLVAEDLNDMPYATKVIKESLRMASIVPWFPRLARQDCQIEGYKIKKGWNINVDARSIHLDPKVYKEPTKFNPSRFDDEG
ISSCFLGFGMGGRTCLGINLAKSMLLVFLHRLLTSYKWKLVDSDSSIEKWAMFSRLKNGCPIHLTRITTP