Lus10024311 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10024311 pacid=23174925 polypeptide=Lus10024311 locus=Lus10024311.g ID=Lus10024311.BGIv1.0 annot-version=v1.0
ATGGGTGAAGGTGATTGTGATTTGCCGAATTCCGTGCTTGTTGAAGCTAACCTAGGGTTGCAGGAAGGTTTGCCAGCGGGAGTGGCTAGACCTAGGGCTG
TGAAGATGGGGAGTGAGGACAAGAAGAAGCCTGGGTTAAAGAAGAGATTTAAGTCAATGGTGGAGAGTGGTGTCGATTCTGATGACGATGAACCGATTGG
GAGTTTGTTGAAGCTGAAGAAGCCTAGGAATCCTAAGAAGGGTAAGGCAGTGTCCGGCAAGGCTTCAAAGGCTCGGAAAGTAATTGCTAAAGATGTGGAA
TTTGGTGGGATGGATGATACATTAGCTAGTTTGAGGAAGAAATTGAAAGGTCCTAAGAAAGACATTGGATCTGTTGCAGCTAGGTGTTCAGATGTTGTTG
TGGAGAAAGGGGTTGGAAACAAGCGCAAGAAGAAAGCAAAACTTGGCGAAGATGCAATGAAATTAGAGTGTGGCGAAGGTAGCAAAGCTATAGGACCTAC
GGATGTTTGCTTCCAATTTAAGAAACGTGAAGGCACATTGGGTGGGGGAACTTCTGTGCATTCTTCAGATGTGAAACTGGTTGAGTCTCAATCGGTTTCT
GCAGCACAGTCTGGCTCAGATCTGCATTACGAACCGAACAGAAGAACCCAAATTTTAGTGAATGGGATAAGCCCTACTTATCGAAGTAGGTTAGATTCTG
CGTCTGAACAATGTAGGACAGTGAAAAACCAGATGCATCAAGATGGTGTTTTTTATAGGATCAAAGATGAAAAAAGGTTCGATTCTGGGGTCTTAGATTG
TGTGGAAAGAGCTGAGGCAGATCCCTGTTGCTCTCACAAAATTTGCGATGATAATTGCTCTTTTTCGGAACAGAAAGGTAGTTCCAACAATAAAACATTT
GAAGATGGTGGACAGCTTTCCAGAGACAGCATATTGGTTCCTGCAGTCCATGGAGTTGCTTATCCCAAATCTTCTTCTATGCAGAATACGACCACATGTA
GTATTGCTGAATTCAATTCCAAGAGTAAGTGCACAAAGACATTGGATGAGAAATTTGAGAATGCCATCACTGAGCCTGTAGTAAATGGTTCTGGTGAAAG
AGAAATTCTATCAACGCTTGATGACAATGTGTTGTTGAATCATCCAAGTGATGTCTCCCCCGATAGTATTCGGGCGAAGGAAACTGTTGAACTGGGGAGT
CTGCCTGTTGAAGTCAAGTGTGAAAAGGATTCTTTCAGCCAACACCCATATACTGCCCAGCCTTCTCGTTCCCATGTAGAAGACTCCAAATTCGTGTTAG
CAGATGTTGAGGAACCTTTTAATCATAATGATACCTTACCTGATGGCCATGATATAGTTTATGTTTCTCCTAAAAGAGAAGCAGCTAAAACACCCGATGG
TAGCTCAACCCCGATTGGTGAAACAGCCCGTGAAGTCCAGAAATCAGAATCTAATCTCCTGATGAGCTGCACAGAGAATTCTCTGGATACAGATGTTGAT
CCACACAACTCCACTGCATCCATTCAAAAGGGAAGCCCCGGTTTGTTACAAAATCAACTTTCTCACAATGAAACCAAGCAGCATTCTGCCCCAGGCCGTG
AACATCATTCCATTAATGAAGGAGTTGATGGAGGCTCTCCTGGATCTTTAGCAGCAGAGGAAAATTGCAGCTTTCAGGACGATACAGCATCTGTTCCTGA
TTCTGAAATCAAAGATGGAAAATTATCAGCTAAGCAACGTGCTGTGCGTCAAGCAAAAAAGCGGAGGCATGAAGACATGGCTTATGAAGGGGATGTAGAT
TGGGAAGTTCTAATAAATGGGCAGAGATTTCTGGATGAGGAAATTTTGGACAATTCGAGAACATACAGAATCAGAGAAAAGTTCGATTCCTCATCAGTAA
GTGTTTCAGAGGTTGAGAATGATGGAGCTGCTGCAGTGTCTGCTGGACTGAAATCCTACGCTGCTGGTCCAGTTGAGAAAATAAAATTCAAAGATATCTT
GAAACGCAAGTCTGGTATCCAAGAATATCTAGAATGCAGGAATAAGATCTTAAGTCTCTGGAGCAAGGATGTTACTCGTATATTGCCACTTTCTGAATGT
GGTGTCACCGATATGCCTTCCGAGGCTGAATCTCAAGGTGCTTCACTAATTAGAGAAGTGTATTCGTATCTTGACCAGAGGGGATATATAAATGTTGGGA
TTGCCTATGAGAAAAAGAAAGCCCAACCAGGTGCGAAGCATAATTATACGCTCTGTGGAGGGAAAGGTCTTGGTGGGAGTCCTAGCACCTCAGTTGCTGA
AATGGATGACGGAGTCTCCTTTATCCTTGGTCAGGTCAAGGGTCGTAAGACTTCAATAGAGGTTAAGGATAGAGTCAGGAATGATAGCGAGAACCAAGGA
TCAGAAAGAGAGCTTGTTACATCTAAGATAAGGTTATTTAATGTCCCAAAACCTGATGAGCTTCTATCTGATGATATCCAGCAAAACATTACTGGCGAAC
CCAGATCATCTGATGGTGTTGTCTCTGCTATAACTCCTCAGTTAAGAGATGACCAGCAAAGTGGGCAATCTTCTCAAAATGATATAGAGAGAGGCCATTA
TATGCCGTGTGATTCTCAAAATAAAAAGAGAATCTTAGTTATTGTTGTTGGTCCTGCTGGATTAACAGCTGCACGGCATCTACAGCGGCAGGGTTTCCAT
GTTACTGTAATTGAGGCACGAAGTAGGATAGGCGGCCGTGTCCATACGGATCGGTCATCCTTTTCAGTTCCAGTAGATCTTGGTGCCAGCATTATTACTG
GTGTTGAGGCGGATGTTGCAACTGAGAGGAGACCAGACCCTTCCTCTTTAATCTGTGCTCAGCTGGGCCTTGAGTTGACTGTTCTAAACAGCGACTGTCC
ACTTTATGATATTGTGACGGGAGAAAAAGTGCCTACTGACCTGGATGAAGCATTGGAGGCAGAATTCAACAGTCTTCTTGATGACATGGTACCACTCATC
TCACAGAAGGGGGAACTTGCTCTGAAGATGTCTCTGGAAGATGGATTAGAATATGCCCTTATGAAGCGGCGCTTAGCACATCCGAGGTCAGATATGGAAG
GAATTGTATTGCAAGACTTTTCATGTGTTTCCACACAATGTAGTGCTGAAGAAGGGGCTGCCCCGAAAAGTTCTAGTGAAGAGATTTTAACTGCTTCTGA
AAGGAGAGTTATGGATTGGCATTTTGCTAATTTGGAGTATGGCTGTGCCACTTTGCTGAAGCAAGTGTCACTTTCGCATTGGAATCAGGACGATGTATAT
GGAGGATTTGGAGGAGCTCATTGCATGATTAAGGGGGGTTACAGTAATGTTGTGGAATCTCTTGCCGAAGGGCTGGTTATTCAACTGAACCATATAGTCA
CTGAGATCTCATATGGTTCCATGGACACGGGTACAAATGATCCAAAAGTAAAAGTTTCCACATCAAATGGCACCGAGTTTAGTGGAGATGCTGTTCTTGT
TACAGTGCCTCTTGGGTGTTTGAAAGCAGAAAGCATAAAGTTTTCCCCTTCCTTACCTCAATGGAAACAGTCTTCCATTCAGAGACTTGGTTATGGAGTC
CTGAATAAAGTAGTCCTGGAATTTGAACGAGTTTTCTGGGATGATTCTGTGGATTACTTTGGGGCGACAGCAGAGGACACTGACAGTAGGGGGCACTGTT
TTATGTTCTGGAATGTACGGAAAACTGTGGGAGCACCTGTTCTTATTGCCTTAGTTGTTGGCAAGGCTGCTATTGATGGCCAAAGTACATGCTCATCAGA
TCACGTAAGTCATGCGGTGAAGACTCTTCGCAAACTTTTTGGAGTGGATTTAGTGCCAGATCCAGTTGCTTCAATGGTGACTGATTGGGGTAGAGATCCT
TTCAGTTATGGTGCTTACTCTTATGTCGCCATAGGTTCATCTGGAGAAGACTATGATATATTGGGCAGACCTGTTGAGAACTGCTTATTTTTTGCTGGTG
AAGCTACCTGCAAGGAGTATCCTGATACAGTTGGTGGTGCTATGATGAGTGGCCTTCGGGAGGCAGTACGGATAATTGACATAATAAGTACTGGAAATGA
TTACACTGCTGAAGCAGAGGCTCTGGAGGCTGCAAAGAGACGGTTGGAACTTGAAAGAGATGAAGTTAGAGACATAATGAAGAGGCTTGAAGCTATGGAG
CTATATACTATCATCTCCAAGAGTTCGGTAGCTTTACTACAGGAGATGTTTTTTGGTGCAAAAACCACTGCAGGAAGACTGCACCTTGCCAAGAAGTTGT
TGAGTCTTCCTGCTGAAACCTTGAAGTCCTTTGCTGGTACCAAAAAAGGGTTATCGACCCTCAACTCCTGGATACTGGACTCGATGGGGAAGGATGGTAC
TCAGCTTTTGCGTCATTGTGTTCGCATTCTTGTGCTGGTATCAACAGACTTACTTGCAGTGCGTTTGTCAGGCATTGGGAAGACAGTCAAAGAAAAAGTT
TGTGTGCATACCAGTCGTGATATTCGTGCAATAGCAAGTCAGCTCGTTAGCGTCTGGCTTGAAGTGTTCCGCAGGGAGAAAGCTTCAAATCGTTTGAAGT
TATCTCGACAAGCCACTGCACTTGATTCTTCAAGGAGAAAGTCCATTTTTAATCTGCTTACAGGGAAGCCACCTCTAAGTAGTCATCTTAATGGTCCGGA
AAATAGGGGGGGCTTGCACTTACCTGCATCCTCTGCTGCTAGCCCATTGTTATCGGATACTAACATGAAGAATGTGAATGAAGAAGAAAAGGCGCCATTA
GCTGCAGCTGAGGCTGCACAAGCTGCAGCTCGTGCAGCTGCTGAGGCTTATGCTAAGTGCAACACGATGGTGCACCTTCCTAAGATACCATCATTTCACA
AGTTTGCAAAACGGGAACAATATCCACCTATGGATGAATATGATTGTAGGAAAAAATGGTCTAGTGGGGTCTTTGGGGGACAAGACTGTACATCTGAGAT
AGACTCGAGGAACTGCAGAGTTAGGAACTGGTCAGTTGATTTTTCCGCTACTTGTGAGAACTTCAGTAATTCCAGATTGTCGATCGATAACTTATCTCAT
AGAAGCCATTCAAACGAAATAGCCTCCCGTATGAACTTGGGAGAGCAGTCTGGAGAAAGTGCGGCTACTGGCAGCAGTCTGTATACAAAAGCATGGGTCG
ATACAGCCGATGTTGGTGCTGTCAAGGGTTATCAAGCCATTGAGAGATGGCAATCTCAAGCAGCTGCTGCTGATTCAGAATTTTACCATCCTGATTTGCA
TATAAACGATGAAGAGCATTCAAATACCAGTTCAAGACCACCTGCCTGGAGGAATGATAGACAAGCAAATGAGAGCTCTGTCTCACAAGTTACTGTAAAC
AAGGAGACTTTAAAGAGCCACTCTCGTAGTGCTGACCGGATTAAACAGGCGGTTGTGGATTTTGTTGCATCATTACTCATGCCTGTTTACAAAGCACACA
AAATTGATAGGGATGGATATAAATCTATAATGAAGAAATGTGCCACAAAGGTGATGGAGCGGGCTACTGATGCCGAGAAAGCAATGTCTGTTTCTGAATT
TCTTGATTCCAAACGCAAGAATAAGATTCGGGCATTTGTTGATAAATTGATGGAGAGGCAAGTGGAAACAGAGCCAGCTGATAAACCGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10024311 pacid=23174925 polypeptide=Lus10024311 locus=Lus10024311.g ID=Lus10024311.BGIv1.0 annot-version=v1.0
MGEGDCDLPNSVLVEANLGLQEGLPAGVARPRAVKMGSEDKKKPGLKKRFKSMVESGVDSDDDEPIGSLLKLKKPRNPKKGKAVSGKASKARKVIAKDVE
FGGMDDTLASLRKKLKGPKKDIGSVAARCSDVVVEKGVGNKRKKKAKLGEDAMKLECGEGSKAIGPTDVCFQFKKREGTLGGGTSVHSSDVKLVESQSVS
AAQSGSDLHYEPNRRTQILVNGISPTYRSRLDSASEQCRTVKNQMHQDGVFYRIKDEKRFDSGVLDCVERAEADPCCSHKICDDNCSFSEQKGSSNNKTF
EDGGQLSRDSILVPAVHGVAYPKSSSMQNTTTCSIAEFNSKSKCTKTLDEKFENAITEPVVNGSGEREILSTLDDNVLLNHPSDVSPDSIRAKETVELGS
LPVEVKCEKDSFSQHPYTAQPSRSHVEDSKFVLADVEEPFNHNDTLPDGHDIVYVSPKREAAKTPDGSSTPIGETAREVQKSESNLLMSCTENSLDTDVD
PHNSTASIQKGSPGLLQNQLSHNETKQHSAPGREHHSINEGVDGGSPGSLAAEENCSFQDDTASVPDSEIKDGKLSAKQRAVRQAKKRRHEDMAYEGDVD
WEVLINGQRFLDEEILDNSRTYRIREKFDSSSVSVSEVENDGAAAVSAGLKSYAAGPVEKIKFKDILKRKSGIQEYLECRNKILSLWSKDVTRILPLSEC
GVTDMPSEAESQGASLIREVYSYLDQRGYINVGIAYEKKKAQPGAKHNYTLCGGKGLGGSPSTSVAEMDDGVSFILGQVKGRKTSIEVKDRVRNDSENQG
SERELVTSKIRLFNVPKPDELLSDDIQQNITGEPRSSDGVVSAITPQLRDDQQSGQSSQNDIERGHYMPCDSQNKKRILVIVVGPAGLTAARHLQRQGFH
VTVIEARSRIGGRVHTDRSSFSVPVDLGASIITGVEADVATERRPDPSSLICAQLGLELTVLNSDCPLYDIVTGEKVPTDLDEALEAEFNSLLDDMVPLI
SQKGELALKMSLEDGLEYALMKRRLAHPRSDMEGIVLQDFSCVSTQCSAEEGAAPKSSSEEILTASERRVMDWHFANLEYGCATLLKQVSLSHWNQDDVY
GGFGGAHCMIKGGYSNVVESLAEGLVIQLNHIVTEISYGSMDTGTNDPKVKVSTSNGTEFSGDAVLVTVPLGCLKAESIKFSPSLPQWKQSSIQRLGYGV
LNKVVLEFERVFWDDSVDYFGATAEDTDSRGHCFMFWNVRKTVGAPVLIALVVGKAAIDGQSTCSSDHVSHAVKTLRKLFGVDLVPDPVASMVTDWGRDP
FSYGAYSYVAIGSSGEDYDILGRPVENCLFFAGEATCKEYPDTVGGAMMSGLREAVRIIDIISTGNDYTAEAEALEAAKRRLELERDEVRDIMKRLEAME
LYTIISKSSVALLQEMFFGAKTTAGRLHLAKKLLSLPAETLKSFAGTKKGLSTLNSWILDSMGKDGTQLLRHCVRILVLVSTDLLAVRLSGIGKTVKEKV
CVHTSRDIRAIASQLVSVWLEVFRREKASNRLKLSRQATALDSSRRKSIFNLLTGKPPLSSHLNGPENRGGLHLPASSAASPLLSDTNMKNVNEEEKAPL
AAAEAAQAAARAAAEAYAKCNTMVHLPKIPSFHKFAKREQYPPMDEYDCRKKWSSGVFGGQDCTSEIDSRNCRVRNWSVDFSATCENFSNSRLSIDNLSH
RSHSNEIASRMNLGEQSGESAATGSSLYTKAWVDTADVGAVKGYQAIERWQSQAAAADSEFYHPDLHINDEEHSNTSSRPPAWRNDRQANESSVSQVTVN
KETLKSHSRSADRIKQAVVDFVASLLMPVYKAHKIDRDGYKSIMKKCATKVMERATDAEKAMSVSEFLDSKRKNKIRAFVDKLMERQVETEPADKP
|
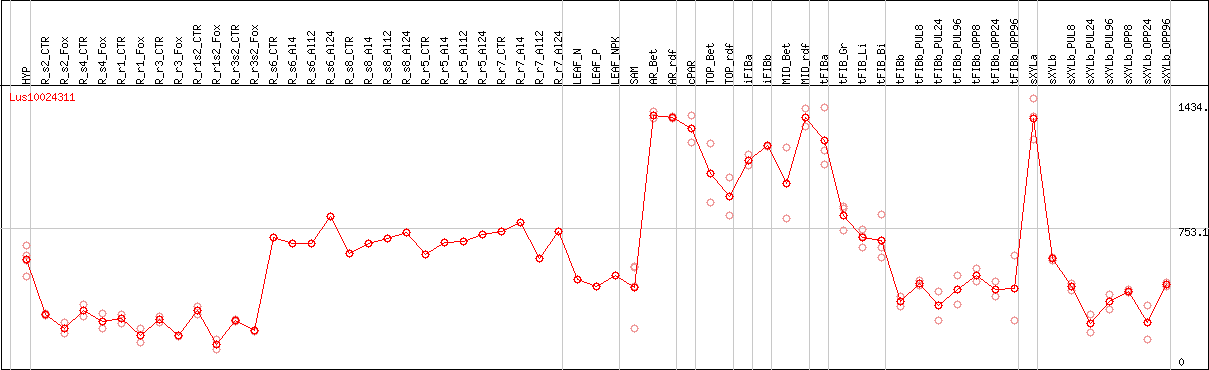
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024311 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
