External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G37660 416 / 4e-147
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G02240 388 / 5e-137
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G31530 86 / 6e-19
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G34460 71 / 5e-14
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G18890 68 / 2e-12
AtTic62
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G16720 54 / 7e-08
HCF173
high chlorophyll fluorescence phenotype 173 (.1)
AT1G15950 45 / 3e-05
IRX4, ATCCR1, CCR1
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010846
498 / 2e-179
AT2G37660 451 / 2e-160
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10022420
82 / 3e-17
AT4G31530 415 / 4e-146
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10009715
79 / 1e-16
AT4G31530 408 / 1e-144
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10040884
71 / 7e-14
AT2G34460 401 / 7e-142
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10004936
69 / 5e-13
AT2G34460 399 / 4e-141
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10042346
68 / 3e-12
AT3G18890 483 / 2e-164
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026321
66 / 1e-11
AT3G18890 479 / 6e-163
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10033322
55 / 4e-08
AT1G16720 874 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Lus10034781
55 / 5e-08
AT1G16720 870 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G086800
436 / 6e-155
AT2G37660 450 / 2e-160
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G253900
71 / 1e-13
AT4G31530 399 / 1e-139
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G150300
70 / 4e-13
AT3G18890 481 / 1e-164
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.T124404
69 / 8e-13
AT3G18890 541 / 0.0
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G111323
69 / 8e-13
AT3G18890 541 / 0.0
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.014G000400
55 / 3e-08
AT1G16720 863 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Potri.007G003500
53 / 1e-07
AT1G16720 852 / 0.0
high chlorophyll fluorescence phenotype 173 (.1)
Potri.003G063800
49 / 3e-06
AT1G72640 348 / 1e-120
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.001G045600
44 / 3e-05
AT1G15950 227 / 4e-75
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
Potri.004G183100
43 / 0.0002
AT4G35250 650 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01073
3Beta_HSD
3-beta hydroxysteroid dehydrogenase/isomerase family
Representative CDS sequence
>Lus10024383 pacid=23174878 polypeptide=Lus10024383 locus=Lus10024383.g ID=Lus10024383.BGIv1.0 annot-version=v1.0
ATGACGTGTCTCCCTTTGATCTCCTTCTCTGCTAAACTCGGTGGCGTGTCCTCATTACTGCCTAGTTCCAGTTGTTGCTCTACACTTCCAGAGTCTTCCG
CACTTCCCTTCTCGCCCTCTCTTTCTGCTTCTTCGACATCGCGGACTACAGGCAGCAGCAGAAGGGCGAGATTCGGTGTTGCGTTTGCATCAATGGCCCC
CAAGACCGTCCTCGTCACCGGAGCTGGTGGCAGAACCGGTTCAATAGTGTACAAGAAACTGAAGGAGAGGTCGGATCATTACACTGCAAGGGGTTTGGTT
CGAACTCCAGAGAGTAAGGAGAAAGTTGGGGGATCTGATGATGTGTACGTTGGAGACGTAAGAGACGCTGAGAGCATTGTTCCAGCTATTCAAGGTATTG
ATGCACTCGTGATACTGACAAGCGCAGTGCCTAAGATGAAACCTGGGTTTGATCCCTCCAAAGGTGAAAGGCCTGAATTTTATTTTGAAGAAGGTTCCTT
TCCTGAACAGGTCGACTGGATTGGACAGAAGAACCAAATAGATGCTGCCAAGGCTGCTGGAGTGAAGCAAATTGTGTTGGTTGGTTCTATGGGTGGAACC
AACCTAGATCACCCATTGAACAAATTGGGCAATGGGAACATACTGGTCTGGAAGAGAAAGGCTGAGCAATATCTGGCTGATTCTGGTCTTCCTTACACAA
TCATAAGGGCTGGAGGGCTGCAAGACAAAGAAGGAGGGATTCGTGAGCTTCTGATCGGGAAGGATGACGAGCTTCTTCAGACAGAGACAAGGACCATAGC
CAGGCCTGATGTTGCTGAAGTGTGCCTTCAGGCTCTCCAGTACGATGAAGCTCAGTTCAAGGCATTTGATTTGGCCTCGAAGCCGGAAGGAACAGGCACC
CCAACCACCGACTTCAAGGCATTGTTCTCTCAGATCACCACCCGGTTTTGA
AA sequence
>Lus10024383 pacid=23174878 polypeptide=Lus10024383 locus=Lus10024383.g ID=Lus10024383.BGIv1.0 annot-version=v1.0
MTCLPLISFSAKLGGVSSLLPSSSCCSTLPESSALPFSPSLSASSTSRTTGSSRRARFGVAFASMAPKTVLVTGAGGRTGSIVYKKLKERSDHYTARGLV
RTPESKEKVGGSDDVYVGDVRDAESIVPAIQGIDALVILTSAVPKMKPGFDPSKGERPEFYFEEGSFPEQVDWIGQKNQIDAAKAAGVKQIVLVGSMGGT
NLDHPLNKLGNGNILVWKRKAEQYLADSGLPYTIIRAGGLQDKEGGIRELLIGKDDELLQTETRTIARPDVAEVCLQALQYDEAQFKAFDLASKPEGTGT
PTTDFKALFSQITTRF
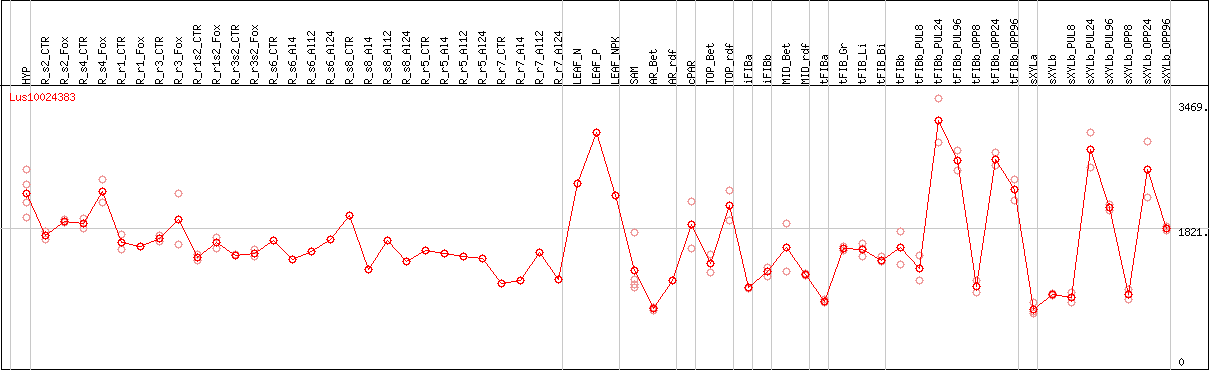
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024383 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.