External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G40860 344 / 8e-115
protein kinase family protein / protein phosphatase 2C ( PP2C) family protein (.1)
AT5G50000 89 / 5e-20
Protein kinase superfamily protein (.1)
AT4G23050 89 / 1e-19
PAS domain-containing protein tyrosine kinase family protein (.1.2)
AT5G49470 87 / 3e-19
PAS domain-containing protein tyrosine kinase family protein (.1.2.3.4)
AT3G01490 86 / 6e-19
Protein kinase superfamily protein (.1)
AT1G67890 86 / 1e-18
PAS domain-containing protein tyrosine kinase family protein (.1.2)
AT1G62400 85 / 1e-18
HT1
high leaf temperature 1, Protein kinase superfamily protein (.1)
AT4G24480 85 / 3e-18
Protein kinase superfamily protein (.1)
AT3G46930 84 / 3e-18
Protein kinase superfamily protein (.1)
AT5G58950 84 / 5e-18
Protein kinase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002993
494 / 2e-173
AT2G40860 899 / 0.0
protein kinase family protein / protein phosphatase 2C ( PP2C) family protein (.1)
Lus10026461
92 / 8e-21
AT4G35780 179 / 7e-51
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Lus10028590
92 / 8e-21
AT4G35780 506 / 9e-175
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Lus10022227
92 / 8e-21
AT3G01490 665 / 0.0
Protein kinase superfamily protein (.1)
Lus10025010
92 / 8e-21
AT4G35780 178 / 1e-49
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Lus10002374
88 / 3e-20
AT1G62400 312 / 7e-107
high leaf temperature 1, Protein kinase superfamily protein (.1)
Lus10000237
88 / 3e-20
AT1G62400 313 / 3e-107
high leaf temperature 1, Protein kinase superfamily protein (.1)
Lus10012517
90 / 6e-20
AT4G35780 754 / 0.0
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Lus10002368
89 / 9e-20
AT5G50000 659 / 0.0
Protein kinase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G030800
298 / 1e-96
AT2G40860 921 / 0.0
protein kinase family protein / protein phosphatase 2C ( PP2C) family protein (.1)
Potri.005G190200
99 / 6e-23
AT4G38470 500 / 2e-172
serine/threonine/tyrosine kinase 46, ACT-like protein tyrosine kinase family protein (.1)
Potri.002G070000
95 / 9e-22
AT4G38470 498 / 1e-171
serine/threonine/tyrosine kinase 46, ACT-like protein tyrosine kinase family protein (.1)
Potri.005G246500
94 / 1e-21
AT4G38470 645 / 0.0
serine/threonine/tyrosine kinase 46, ACT-like protein tyrosine kinase family protein (.1)
Potri.006G222600
94 / 2e-21
AT4G35780 175 / 1e-48
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Potri.015G074400
90 / 3e-20
AT5G50000 654 / 0.0
Protein kinase superfamily protein (.1)
Potri.007G060600
90 / 6e-20
AT4G35780 793 / 0.0
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Potri.004G179100
89 / 7e-20
AT4G35780 612 / 0.0
serine/threonine/tyrosine kinase 17, ACT-like protein tyrosine kinase family protein (.1)
Potri.002G015400
89 / 9e-20
AT4G38470 650 / 0.0
serine/threonine/tyrosine kinase 46, ACT-like protein tyrosine kinase family protein (.1)
Potri.001G351300
88 / 1e-19
AT3G01490 664 / 0.0
Protein kinase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10024475 pacid=23170258 polypeptide=Lus10024475 locus=Lus10024474.g ID=Lus10024475.BGIv1.0 annot-version=v1.0
ATGGGTTTGCAGATCGCGGAGCCAAACTCTTGCATTAGAGGATGCTGCAGCAGCGATTCAATCCCGCTTCATCTACCAACTTCTTCTTACACTCTCTTAT
CTCCGATTGCCAGAGGAGCTGAGAGCGTGGTATATGAAGCAATCTTGAACGATGTTAAAGTTGCTGTGAAGAAACCTATCTTGTCTACTTCTGATCACAT
TGATAAGTTCCACAAAGAGCTGCAACTATTGTGTACATTTGACCTTCCAGGGATATCGAGGCATGTGAAACCTGCAAACATGCTTCTAGATGAGAACCTT
TATCCCCACCTTGCCGACTTTGGCCTGGCAGAGTACAAGAATAACCTTAAGCAAGTCACTTTAGACAACTGGAGATCTTCGGGGAAGCCTACTGGTGGGT
TTCATAAAAAGAATATGGTTGGTACGCTTATTTATATGGCCCCTGAAGTATTGAGGAAGGAGGTTCAGACTGAGAAATCAGATGTTTATAGCTTTGGGAT
ATCCATCAACGAGCTTCTTACTGGTGTCGTCCCGTATACTGATCTTCGTGCAGAGGCTCAAGCCCACACTGTCTTGGAAATGAACTATACGGAGCAGCAA
CTTACAGCCGCTGTGGTATCTGGTAAATTGAGACCTGCTCTTGCAATTTCTGAGTCAGGTGCATCACCAAGTCTTCTTTTTTTAATACAGAGATGCTGGG
ATGACAATCCTGATCGTAGACCTTCTTTTGATGAAATAGTTGTAGAACTAGACACAATGATAGGTCAGGAACTGGTCCAAGCGAACCATGTCTGTGATAT
GGCTTCTGATTCTACCAGGGATAAAGCGAAAGGCACCATAATGAACCGAAAGGCACCATAA
AA sequence
>Lus10024475 pacid=23170258 polypeptide=Lus10024475 locus=Lus10024474.g ID=Lus10024475.BGIv1.0 annot-version=v1.0
MGLQIAEPNSCIRGCCSSDSIPLHLPTSSYTLLSPIARGAESVVYEAILNDVKVAVKKPILSTSDHIDKFHKELQLLCTFDLPGISRHVKPANMLLDENL
YPHLADFGLAEYKNNLKQVTLDNWRSSGKPTGGFHKKNMVGTLIYMAPEVLRKEVQTEKSDVYSFGISINELLTGVVPYTDLRAEAQAHTVLEMNYTEQQ
LTAAVVSGKLRPALAISESGASPSLLFLIQRCWDDNPDRRPSFDEIVVELDTMIGQELVQANHVCDMASDSTRDKAKGTIMNRKAP
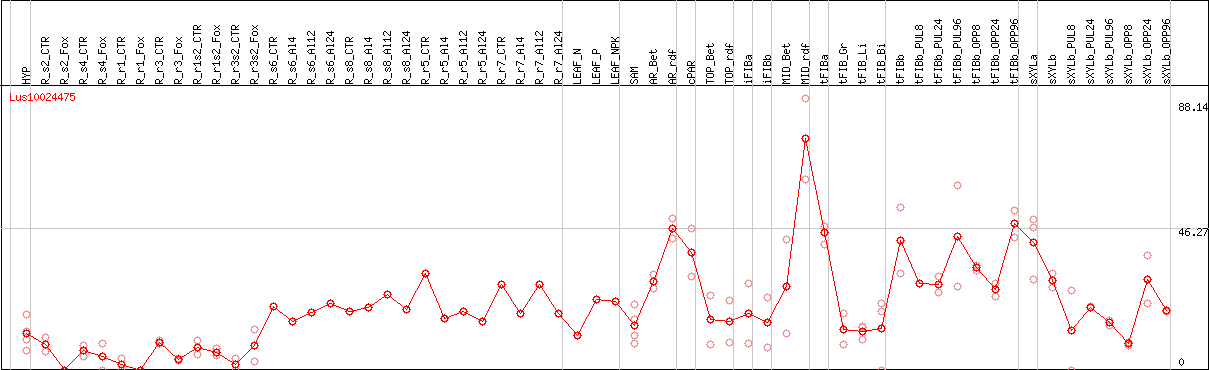
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024475 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.