External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G03080 443 / 2e-151
BSL1
BRI1 suppressor 1 (BSU1)-like 1 (.1)
AT2G27210 407 / 3e-136
BSL3
BRI1 suppressor 1 (BSU1)-like 3 (.1), BRI1 suppressor 1 (BSU1)-like 3 (.2)
AT1G08420 401 / 9e-134
BSL2
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
AT1G03445 305 / 1e-98
BSU1
BRI1 SUPPRESSOR 1, Serine/threonine protein phosphatase family protein (.1)
AT5G59160 182 / 4e-56
PPO, TOPP2
PROTOPORPHYRINOGEN OXIDASE, type one serine/threonine protein phosphatase 2 (.1.2.3)
AT3G46820 179 / 3e-55
TOPP5
type one serine/threonine protein phosphatase 5 (.1)
AT2G39840 173 / 1e-52
TOPP4
type one serine/threonine protein phosphatase 4 (.1)
AT5G27840 172 / 2e-52
TOPP8
Calcineurin-like metallo-phosphoesterase superfamily protein (.1.2)
AT2G29400 169 / 3e-51
PP1-AT, TOPP1
type one protein phosphatase 1 (.1)
AT5G43380 169 / 6e-51
TOPP6
type one serine/threonine protein phosphatase 6 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009783
460 / 1e-158
AT2G27210 1281 / 0.0
BRI1 suppressor 1 (BSU1)-like 3 (.1), BRI1 suppressor 1 (BSU1)-like 3 (.2)
Lus10008369
410 / 6e-137
AT1G08420 1747 / 0.0
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
Lus10004100
406 / 1e-135
AT1G08420 1686 / 0.0
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
Lus10013357
406 / 2e-135
AT1G08420 1713 / 0.0
BRI1 suppressor 1 (BSU1)-like 2 (.1), BRI1 suppressor 1 (BSU1)-like 2 (.2)
Lus10016489
180 / 2e-55
AT5G59160 570 / 0.0
PROTOPORPHYRINOGEN OXIDASE, type one serine/threonine protein phosphatase 2 (.1.2.3)
Lus10011827
180 / 7e-55
AT2G39840 513 / 0.0
type one serine/threonine protein phosphatase 4 (.1)
Lus10040259
179 / 9e-55
AT2G39840 590 / 0.0
type one serine/threonine protein phosphatase 4 (.1)
Lus10001555
176 / 1e-54
AT5G59160 470 / 2e-169
PROTOPORPHYRINOGEN OXIDASE, type one serine/threonine protein phosphatase 2 (.1.2.3)
Lus10004692
178 / 2e-54
AT2G39840 596 / 0.0
type one serine/threonine protein phosphatase 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0163
Calcineurin
PF00149
Metallophos
Calcineurin-like phosphoesterase
Representative CDS sequence
>Lus10024772 pacid=23159524 polypeptide=Lus10024772 locus=Lus10024772.g ID=Lus10024772.BGIv1.0 annot-version=v1.0
ATGCTCTCTCAGATCGAGTATCCAGAAAATGTTCACCTGATACGAGGAAACCATGAAGCTGCTGATATTAATGCACTCTTTGGTTTTCGCATTGAATGCA
TTGAGAGAATGGGAGAGAGTGATGGGATATGGGCATGGACACGCTTCAATCAGCTCTTCAACTATCTTCCACTTGCAGCACTTGTTGAAAAGAAAATAAT
ATGTATGCATGGGGGAATCGGCAGATCTATTACTTCAGTGGAACAGATTGAAAAACTCGAGAGACCCATAACCATGGATGCAGGTTCCATAATACTGATG
GATCTTCTGTGGTCTGATCCTACAGAAAATGATAGTGTGGAAGGTTTAAGGCCAAATGCGAGAGGCCCTGGTCTTGTCACTTTCGGGCCTGATCGTGTAA
CAGATTTTTGTAAGAAGAACAAGTTGCAGCTGATTATTCGAGCACATGAATGTGTTATGGATGGGTTTGAACGGTTTGCTCAAGGACAGCTGATTACGTT
ATTTTCTGCTACGAATTACTGCGGGACTGCAAACAACGCGGGAGCCATACTAGTGGTTGGACGGGGATTGGTTGTGGTACCAAAGTTAATCCATCCATTG
CCGCCACCTCTTCAATCACCGGAAACATCTCCGGAGCGTATTGCAGAAGAGACATGGATGCAAGAGCTTAACATCCAGAGACCACCAACTCCGACTCGGG
GTCGCCCACAACCTGACCTTGACAGAAACTCACTTGCTTACATATGA
AA sequence
>Lus10024772 pacid=23159524 polypeptide=Lus10024772 locus=Lus10024772.g ID=Lus10024772.BGIv1.0 annot-version=v1.0
MLSQIEYPENVHLIRGNHEAADINALFGFRIECIERMGESDGIWAWTRFNQLFNYLPLAALVEKKIICMHGGIGRSITSVEQIEKLERPITMDAGSIILM
DLLWSDPTENDSVEGLRPNARGPGLVTFGPDRVTDFCKKNKLQLIIRAHECVMDGFERFAQGQLITLFSATNYCGTANNAGAILVVGRGLVVVPKLIHPL
PPPLQSPETSPERIAEETWMQELNIQRPPTPTRGRPQPDLDRNSLAYI
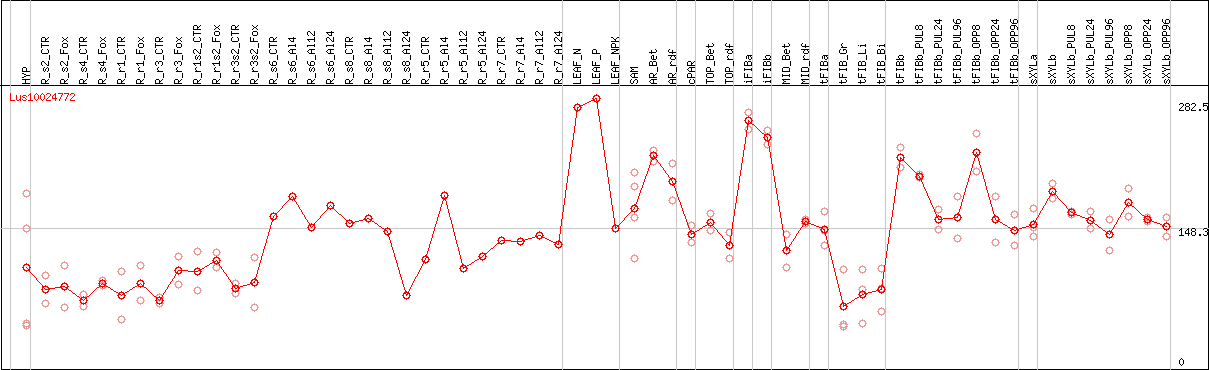
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024772 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.