Lus10024805 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10024805 pacid=23178923 polypeptide=Lus10024805 locus=Lus10024805.g ID=Lus10024805.BGIv1.0 annot-version=v1.0
ATGGATACTCCTCCACCGTCTCCCGGCGCCGGTGCCGGTGCTGGGGGAGAGAATTGCTGTGTAAAAGTAGCCGTACATGTAAGACCCCTAATTGCCGACG
AAAGAGCTCAAGGCTGTAAAGATTGCGTCACTGTCGTCCCTGGAAAACCTCAGGTACAAATAGGGACGCACTCGTTTACCTTCGATCATGTATATGGGAG
CACAGCTTCACCATCCTCTGCTATGTTTGAGGAATGTGTTTCTCCACTTGTTGATGGCTTGTTCCAAGGTTACAATGCAACAGTTCTCGCCTATGGTCAG
ACAGGGTCGGGAAAGACTTATACAATGGGTACTGGTTTCAAAGATGGTTATCAAGCTGGAATAATTCCTCAAGTTATGAGTGTTCTCTTTGGAAAGATCG
ACACTTTGAAGCATCAAACTGAATTCCAGCTTCATGTTTCTTTCATTGAGATTCTGAAAGAGGAAGTGCGAGACTTGCTTGATTCCGCTTTACTGAACAA
AATGGAATCTGGAAATGGGAATGCTGGCAAAGTGCACGTTCCTGGAAAACCGCCGATACAGATTCGTGAATCGTCAAATGGTGTGATCACATTAGCAGGA
TCCACGGAGGTTAGTGTAAGCAGTTTTAAGGAAATGGCTGCTTACCTAGAGCAGGGATCAATGAGCAGGGCAACAGGGAGTACAAACATGAACAACCAAT
CTAGTCGCTCCCATGCCATTTTCACCATCACCCTAGAGCAGATACACAAACTCAATCCAATGTTTCCTGGAGATGGGTTTCCTAATGACAGCATGAATGA
TGAGTATCTTTGTGCTAAATTGCATTTAGTAGATCTTGCTGGATCAGAGAGGGCTAAGAGGACTGGTTCTGATGGTTTGCGTTTCAAGGAAGGCATTCAC
ATAAACAAGGGCCTTCTGGCACTTGGCAATGTTATCAGTGCACTTGGCGATGAGAAAAAGCGTAAAGAAGGTCTTCATGTTCCTTATCGGGACAGTAAAT
TGACCCGCCTTTTGCAGGCGAGTATTCATACCCTAGGCATCCTGGTTTCTCTTCGTGCTTTCGACTCACTTGGCGGTAACAGCAGAACCGTCATGATAGC
TTGCATTAGCCCTGCAGATATAAATGCTGAGGAAACCCTTAATACTCTGAAATATGCAAATCGTGCCCGAAATATTCAAAACAAGCCTGTTGTCAATAGA
GATCCTATGTCCAGTGAGATGCTCAAAATGCGACAACAACTCGAGTATCTTCAGGCAGAACTTTGTGCCCGTGGTGGATCCTCACCTGTTGAAGTTCAGG
TTCTCAAAGAAAGAATTGCTTGGCTAGAAGCTACGAATGAGGACCTTTGTAGAAAGCTTCATGAATACCGCAGCAGGTGTGCTCCTGGGGACCTGCGTGA
AATAGATGCTCAAGATAGTAGTATCTGTTCTATAAAAGGTGAAGCAGTTAGAAGGAGCTTACATAGTCTAGAGGCATCTGACTATCAAATGGTTGAAACA
ATATCAGCTCATACCAAGGAAATTGATGAAGAAGTAGCAAAAGAATGGGAGCACACACTTCTCCAGAACACACTAGACAAGGAGCTACATGAGTTAAACA
GACGCTTGGAAGAGAAAGAGTCAGAGATGAAACTTGTTGGGGGATCTGACACTGCAGCACTTAAGCAACACTTCGGCAAGAAACTCATGGAATTAGAGGA
TGAAAAAAGAGTTGTGCAGCAAGAACGTGATCGTTTGTTAGCCGAAATTGAAAACCTATCGGCTAACTCAGATGGGCAGACACAGAAATTGCAAGATGTG
CATGCCCAGAAACTGAAGACATTAGAAGGGCAGATTCTGGATCTTAAGAAGAAACAGGAGAACCAAGTTCAGCTTTTGAAGCAAAAGCAAAAAAGTGACG
AAGCAGCAAGGAAATTGCAAGAAGAAATACAATCTATAAAAGCTCAAAAGGTTCAACTGCAACACAGAATGAAGCATGAGGCAGAACAATTTCGACAATG
GAAGGCCTCTAGAGAAAAGGAATTGCTTCAGTTACGCAAAGAGGGGAGAAGAAACGAATACGAAAGGCACAAGTTACAAGCCATAAACCAGCGCCAAAAA
ATGGTTCTGCAGAGAAAAACAGAAGAAGCTGCTGTTGCTACCAAAAGGCTGAAGGAGTTACTCGAAGGTCGAAAATCTTCACCACGGGATAGCACAGGAC
TTGCAAATGGACATGTAGCCAATGGGCAGAGTAATGAGAAATCCTTGCAGCGATGGCTTGAGCATGAGCTAGAAGTGATGGTGAATGTCCATGAAGTTCG
GTTTGAATATGAGAAGCAGAGTCAAGTGAGAGCTGCTCTAGGGGAAGAACTTGCTGTGTTGAAGGAAGTGGATGAGCTAGCTTCAAAAGGATTAAGCCCT
CCAAAGGGGAAGAATGGATTTTCCAGGGCTTCTTCCATGTCACCAAATGCCAGAATGGCCAGAATAGCATCGCTGGAGAACATGTTAAACATATCATCCA
ACTCGCTTGTGGCAATGGCTTCGCAACTTTCAGAGGCAGAGGAACGTGAACGTGGTTTGACTACCCGTGGGCGTTGGAACCAATTGCGCTCCATGGGAGA
TGCAAAGAACTTGCTTCAGTATATGTTTAACTCTCTTGGAGATGCAAGGTGCCAATGCTGGGAGAAGGATATGGAAATCAAGGAAATGAAGGAACAATTC
AAAGAACTTGTGAACCTATTACGACAAAGTGAGACTCGAAGAAAGGAAGTCGAGAAGGAGCTGATACATAGAGAGGAAGCTATTGCTATTGCCTTGGCTA
CAGCAGCTTCGGCTGGTCATGAACAAGGGAACTCACCCAATTCACTAAAACACAGTGCTGATGAAATGATGAGCAGTCCCTTGTCTCCGGTGTCTGTGCC
AGCACAGAAGCAGCTGAAATATACGCCAGGGATCGTCAATGCCCCAATCACAGAAGCTGCGGCATTTATAGATCAGTCGAGAAAAATGGTGCCACTTGGG
GACGCATCAATGAGAAAATTAGCAATTGCAGGACAAAATGGGAAGCTATGGAGATGGAAGAGAAGCCACCACCAGTGGCTTTTGCAATTTAAATGGAAAT
GGCAGAAGCCATGGAGGATCTCTGAGACGATCAGGCTTAACGATGTAACAATTTCAAGGACAAGGCCTCGTGTACAGTATCGAGCACGAGTGTCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10024805 pacid=23178923 polypeptide=Lus10024805 locus=Lus10024805.g ID=Lus10024805.BGIv1.0 annot-version=v1.0
MDTPPPSPGAGAGAGGENCCVKVAVHVRPLIADERAQGCKDCVTVVPGKPQVQIGTHSFTFDHVYGSTASPSSAMFEECVSPLVDGLFQGYNATVLAYGQ
TGSGKTYTMGTGFKDGYQAGIIPQVMSVLFGKIDTLKHQTEFQLHVSFIEILKEEVRDLLDSALLNKMESGNGNAGKVHVPGKPPIQIRESSNGVITLAG
STEVSVSSFKEMAAYLEQGSMSRATGSTNMNNQSSRSHAIFTITLEQIHKLNPMFPGDGFPNDSMNDEYLCAKLHLVDLAGSERAKRTGSDGLRFKEGIH
INKGLLALGNVISALGDEKKRKEGLHVPYRDSKLTRLLQASIHTLGILVSLRAFDSLGGNSRTVMIACISPADINAEETLNTLKYANRARNIQNKPVVNR
DPMSSEMLKMRQQLEYLQAELCARGGSSPVEVQVLKERIAWLEATNEDLCRKLHEYRSRCAPGDLREIDAQDSSICSIKGEAVRRSLHSLEASDYQMVET
ISAHTKEIDEEVAKEWEHTLLQNTLDKELHELNRRLEEKESEMKLVGGSDTAALKQHFGKKLMELEDEKRVVQQERDRLLAEIENLSANSDGQTQKLQDV
HAQKLKTLEGQILDLKKKQENQVQLLKQKQKSDEAARKLQEEIQSIKAQKVQLQHRMKHEAEQFRQWKASREKELLQLRKEGRRNEYERHKLQAINQRQK
MVLQRKTEEAAVATKRLKELLEGRKSSPRDSTGLANGHVANGQSNEKSLQRWLEHELEVMVNVHEVRFEYEKQSQVRAALGEELAVLKEVDELASKGLSP
PKGKNGFSRASSMSPNARMARIASLENMLNISSNSLVAMASQLSEAEERERGLTTRGRWNQLRSMGDAKNLLQYMFNSLGDARCQCWEKDMEIKEMKEQF
KELVNLLRQSETRRKEVEKELIHREEAIAIALATAASAGHEQGNSPNSLKHSADEMMSSPLSPVSVPAQKQLKYTPGIVNAPITEAAAFIDQSRKMVPLG
DASMRKLAIAGQNGKLWRWKRSHHQWLLQFKWKWQKPWRISETIRLNDVTISRTRPRVQYRARVS
|
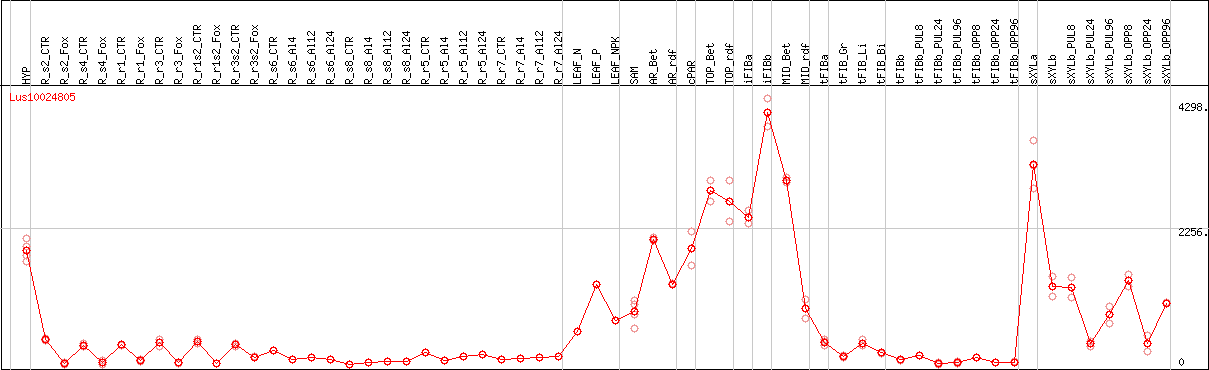
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024805 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
