External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G38940 216 / 3e-69
PHT1;4, ATPT2
ARABIDOPSIS THALIANA PHOSPHATE TRANSPORTER 2, phosphate transporter 1;4 (.1)
AT3G54700 215 / 1e-68
PHT1;7
phosphate transporter 1;7 (.1)
AT5G43360 202 / 7e-64
PHT1;3, ATPT4, PHT3
PHOSPHATE TRANSPORTER 3, phosphate transporter 1;3 (.1)
AT2G32830 202 / 1e-63
PHT1;5, PHT5
PHOSPHATE TRANSPORTER 5, phosphate transporter 1;5 (.1)
AT5G43350 199 / 1e-62
PHT1;1, ATPT1
ARABIDOPSIS THALIANA PHOSPHATE TRANSPORTER 1, phosphate transporter 1;1 (.1)
AT5G43370 182 / 4e-56
PHT1;2, APT1, PHT2
ARABIDOPSIS PHOSPHATE TRANSPORTER 1, phosphate transporter 2 (.1.2)
AT5G43340 179 / 5e-55
PHT1;6, PHT6
PHOSPHATE TRANSPORTER 6, phosphate transporter 1;6 (.1)
AT1G20860 147 / 1e-42
PHT1;8
phosphate transporter 1;8 (.1)
AT1G76430 145 / 4e-42
PHT1;9
phosphate transporter 1;9 (.1)
AT4G08895 125 / 9e-38
inorganic phosphate transporter family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014754
223 / 2e-71
AT2G38940 904 / 0.0
ARABIDOPSIS THALIANA PHOSPHATE TRANSPORTER 2, phosphate transporter 1;4 (.1)
Lus10033886
219 / 5e-70
AT3G54700 919 / 0.0
phosphate transporter 1;7 (.1)
Lus10003560
219 / 6e-70
AT3G54700 919 / 0.0
phosphate transporter 1;7 (.1)
Lus10011826
205 / 1e-64
AT5G43360 868 / 0.0
PHOSPHATE TRANSPORTER 3, phosphate transporter 1;3 (.1)
Lus10021191
196 / 4e-61
AT5G43350 858 / 0.0
ARABIDOPSIS THALIANA PHOSPHATE TRANSPORTER 1, phosphate transporter 1;1 (.1)
Lus10016635
191 / 2e-59
AT3G54700 888 / 0.0
phosphate transporter 1;7 (.1)
Lus10022548
175 / 4e-54
AT3G54700 578 / 0.0
phosphate transporter 1;7 (.1)
Lus10025163
148 / 9e-44
AT1G76430 431 / 5e-148
phosphate transporter 1;9 (.1)
Lus10030506
147 / 1e-42
AT5G43360 665 / 0.0
PHOSPHATE TRANSPORTER 3, phosphate transporter 1;3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0015
MFS
PF00083
Sugar_tr
Sugar (and other) transporter
Representative CDS sequence
>Lus10024884 pacid=23178881 polypeptide=Lus10024884 locus=Lus10024884.g ID=Lus10024884.BGIv1.0 annot-version=v1.0
ATGAACGCGATCGAAGAGGTTTTCCATATCACCAGAGCTCAGACGCTAATCGTGATTTGCAGCACCGTGCCCAGTTACTGGTTCACGGTAGCTTTGATCA
ACAAAATCGGGAGGTTCGCTATCCAATTGATGGGGTTCTTCTTCATGACAGTGTTCATGTTTGCTCTCGCGGTACCTTACAGTCACTGGACTCACAGGGA
GAATCGGATCGGGTTCGTGGTGCTCTACGCACTGACATTCTTCTTCGCGAACTTCGGGCCTAATGCGACAACATTTGTGGTTCCAGCGAAGATATTCCCG
GCCAGGGTGAGGTCTACTTGCCATGGGATTTCGTCGGCTTTGGGGAAAGTGGGGGCGATCGGGGTTCTTGTACTTGGCGCAGAATCAGGATAA
AA sequence
>Lus10024884 pacid=23178881 polypeptide=Lus10024884 locus=Lus10024884.g ID=Lus10024884.BGIv1.0 annot-version=v1.0
MNAIEEVFHITRAQTLIVICSTVPSYWFTVALINKIGRFAIQLMGFFFMTVFMFALAVPYSHWTHRENRIGFVVLYALTFFFANFGPNATTFVVPAKIFP
ARVRSTCHGISSALGKVGAIGVLVLGAESG
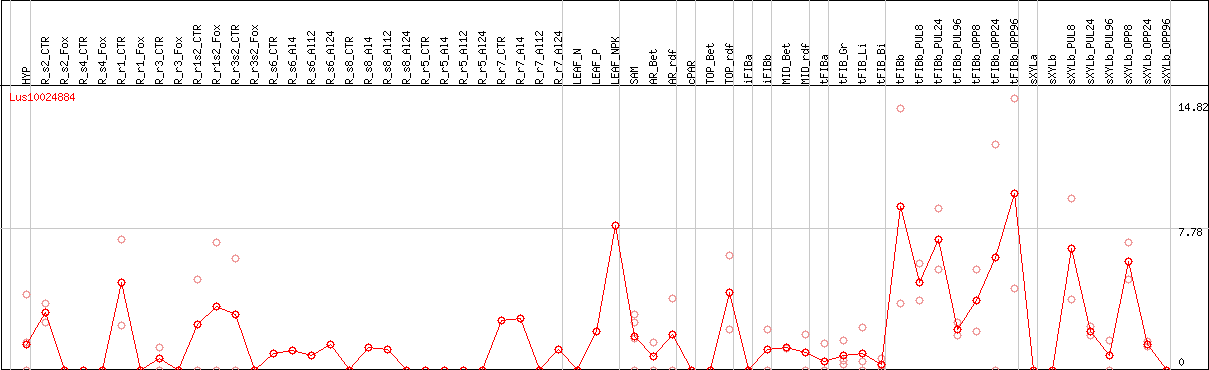
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10024884 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.