External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G21060 110 / 3e-32
ATCSP4, ATGRP2B
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
AT2G17870 101 / 2e-27
ATCSP3
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
AT4G38680 94 / 2e-25
ATCSP2, CSDP2, GRP2
COLD SHOCK DOMAIN PROTEIN 2, ARABIDOPSIS THALIANA COLD SHOCK PROTEIN 2, glycine rich protein 2 (.1)
AT4G36020 93 / 2e-24
CSDP1
cold shock domain protein 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005066
130 / 2e-40
AT2G17870 122 / 1e-35
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
Lus10027839
130 / 2e-40
AT2G21060 110 / 6e-31
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Lus10034487
122 / 4e-37
AT2G21060 110 / 7e-31
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Lus10028073
92 / 8e-25
AT4G36020 115 / 3e-31
cold shock domain protein 1 (.1)
Lus10028449
79 / 2e-19
AT4G36020 158 / 3e-47
cold shock domain protein 1 (.1)
Lus10041902
79 / 2e-19
AT4G36020 159 / 2e-47
cold shock domain protein 1 (.1)
Lus10025623
74 / 2e-18
ND 81 / 8e-20
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G172600
119 / 2e-35
AT2G21060 102 / 3e-27
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Potri.009G132000
112 / 8e-33
AT2G21060 103 / 1e-27
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Potri.005G174300
93 / 4e-25
AT2G17870 92 / 1e-22
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
Potri.002G086900
89 / 1e-23
AT2G17870 89 / 2e-21
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
Potri.007G057100
76 / 5e-18
AT4G36020 127 / 3e-35
cold shock domain protein 1 (.1)
Potri.005G112800
75 / 6e-18
AT2G17870 86 / 1e-19
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0021
OB
PF00313
CSD
'Cold-shock' DNA-binding domain
Representative CDS sequence
>Lus10025058 pacid=23167822 polypeptide=Lus10025058 locus=Lus10025058.g ID=Lus10025058.BGIv1.0 annot-version=v1.0
ATGAGTGCCAGATTGTCGGGCAGGGTGAAGTGGTTCAATGATCAGAAGGGGTACGGCTTCATCACCCCTAACGACGGCAGCGAGGATCTCTTCGTTCACC
AGTCCTCCATCCAGACCGAAGGCTTCCGCAGCCTCGCCGAAGACGAAGAGGTGGAGTACGTCATCGAGAAGTCCAACGATGGCCGCACCAAGGCTATTGA
AGTGACCGGTCCGGATGGAAAGCCCGTCCAAGGCGGAGGACGCGGTGGTGGAGGCGGGTGA
AA sequence
>Lus10025058 pacid=23167822 polypeptide=Lus10025058 locus=Lus10025058.g ID=Lus10025058.BGIv1.0 annot-version=v1.0
MSARLSGRVKWFNDQKGYGFITPNDGSEDLFVHQSSIQTEGFRSLAEDEEVEYVIEKSNDGRTKAIEVTGPDGKPVQGGGRGGGGG
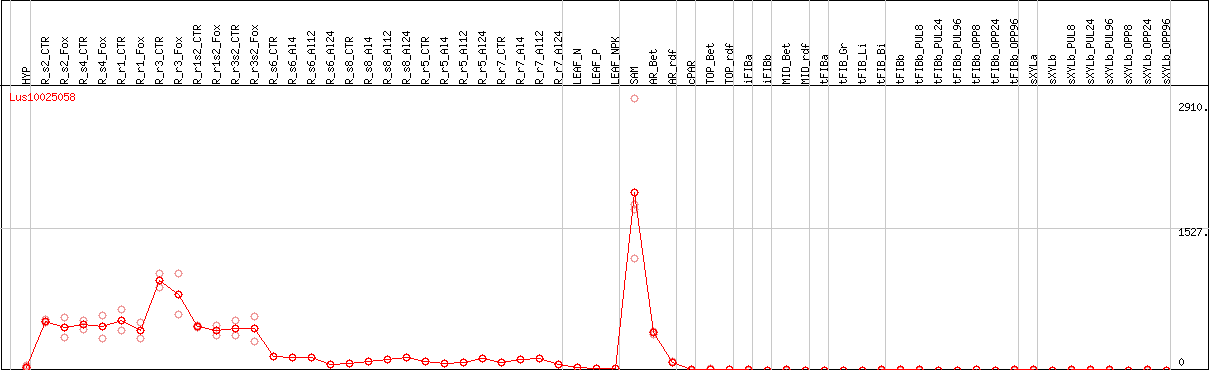
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025058 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.