Lus10025071 [FLAX]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10025071 pacid=23167869 polypeptide=Lus10025071 locus=Lus10025071.g ID=Lus10025071.BGIv1.0 annot-version=v1.0
ATGAGCTCTCAACCTAGAGGCGATCGAAGGGACAGGTCTAGATCATCTTCTCAAACTGCACAGCAGCGTTGGATTCCCCGAGGATCTGCCCCCTCATCAG
TTTCTGTCAACAATAACATACCTCCTGCAACTTTCAATCCAAGTAACGTTGATGGAAGAGGTGAGCAGTCGAATGCTAATCCCAATTCCGGACCCCGTGA
GGCTAGACCTAAAGGAGCCGGCAATGCTTCTCGTAGTTATGTGGCGGGTCGTCCGGTGAATAATCACAGGAGGGACAAGCAAAGGGAGAAGGGTGAAACA
CATGCTGTGAAGGAGTCGGTTGATCCGAACCTGCCTCAACTTGTTCAAGAGATTCGGGAGAAGCTTGCAAAGGGAACTGTCGAGTGCATGATCTGTTATG
ATATGGTGCGAAGGTCTGCTCCTATTTGGTCTTGCTCGAGCTGCTACTCCATTTTTCATCTGCATTGTATCAAGAAATGGGTGAGAGCTCCTACATCGAT
CGACATGTCAGCAGATAAGCATCAGGGATTTAATTGGCGCTGCCCAGGCTGCCAATATGTTCAGCTCACGCCATTGAAGGAGATCAGCTATCTCTGCTTT
TGTGGGAAAAGAACTGATCCACCTTCTGATTTGTATTTGACTCCTCATTCATGTGGAGAGCCATGTGGGAAGCCGCTCGAGAGGCATTTACCAGGTAGCT
TGGAGACTAAGGAGGACTCTTGCCCTCATAACTGTGTGTTGCAGTGCCACCCCGGGCCCTGCCCACCATGTAAATCATTTGCTCCACCATGCTTGTGTCC
TTGCGGGAAGAAAACTATTACCACTAGGTGTTCTGATCGCCAGTCTGTTCTCACGTGTGGTCAACAATGTGATAAGCTTCTTGATTGTCAGCGTCATCGC
TGTGGGAGGATGTGCCATGTCGGACCTTGCGATCCTTGTGAAATCCTTGTCACAGCCTCGTGCTTTTGCAAGAAAAAGCAGGAGGTTGTTTATTGCGGAG
ACATGTCTGTGAAGGGAGAAGTGAACGTTAAAGACGGGGTGTTTTCCTGCAATTCCAGTTGTGGAAAGCTGCTTGCTTGTGGAAATCATGAATGCGATGA
AGTGTGTCATCCAGGTCCTTGTGGCAGTTGTGCTTTGATGCCAGGAAGAGTCAAGTCTTGTTATTGTGGAAAGACGAGCTTGCAGAAGGAAAGGAAGTCT
TGTTTGGATCCTATTCCTAACTGTTCACAGGTTTGCAGCAAACTTCTTCCCTGTGGAATTCACAACTGTGAAGACTTGTGCCATTCTGGTGATTGCCCGC
CTTGTATGGTCAGTGTCTCGCAGAAATGTCAATGTGGATCAACCTCCCGAACTGTGGCATGTTACCAAACAAGTACTGAAAGTGAAAAATTCAAGTGTGA
CAAGCCCTGTGGTCGCAAGAAGAACTGTGGAAGGCATCGGTGCAGTGAACGTTGCTGCCCTCTTTCAAATCCGAATCCTGTTCGGTCTGAGGATTGGGAT
CCACACTTCTGCCAAATGGCGTGTGGGAAGAAGTTGAGGTGTGGTCAACATTCTTGTGAATCTCTGTGTCACAGTGGTCACTGTCCTCCCTGTCTCGAGA
CAATATTCAGTGATTTAACTTGTGCCTGTGGAAGGACTTCAATCCCTCCGCCATTGCCGTGTGGTACACCGGCACCTTCATGTCAGCTCCCATGTTCAGT
TCCTCAACCTTGTGGGCATTTGGCTTCTCATAGCTGCCACTACGGTGACTGCCCTCCTTGTTCAGTCCCTGTTGCAAAGGAATGTGTTGGTGGGCATGTG
GTTCTTAAAAACATACCATGCGGGTCAAAAGACATAAGGTGTAATAAGCTCTGTGGTAAAACTAGGCAGTGTGGTCTACATGCATGTGGGCGAACTTGTC
ATCCAGCACCATGCGACTCTTCTGGTTCAACTAGTGCAGGGGCGGCGAGAGTTTCTTGTGGCCAGACATGTGGGGCGCCCAGAAGAGACTGCAGGCATAC
TTGTGCAGCACTTTGCCATCCTTCTTCTCCTTGCCCAGACGTGAGATGCCAGTTCCCTGTGACGATTAGTTGCTCATGCGGCAGGATAACAGCTAGCGTG
CCATGCGATGCCGGAGGGACCAATGCTGGTTATAGTGCCGATACGGTTTTTGATGCTTCCATAGCACATAAGTTGCCAGCACCGCTTCAACCAGTGGAAT
CAACTGGTTCGAGGATCCCACTTGGCCAACGGAAGCTCACCTGTGATGAGGAATGTGCTAAGTCGGAGCGTAAGAGGGTTCTCGCTGATGCTTTTGACAT
TAGTACTCCGAATTTCGAAGCTCTTCATTTTGGCGAGAACTCGCAGGTGACGGAATTGATTGCTGATTTATATAGGCGTGACCCGAAATGGGTTCTTGCC
GTGGAAGAGAGGTGCAAGTACCTTGTACTTGGGAAGAGCAAGGGAACTGTAGGTGGTCTAAAGGTTCACGTTTTTTGTCCAATGATGAAGGACAAGCGTG
ATGCAGTTAGGCTGATTGCAGAGAGGTGGAAGCTATCGATAAACGTTGCTGGTTGGGAGCCGAAGCGGTTCGTTGTGCTACATGCAACTCCCAAGTCCAG
ACCCCCATCTCGCGTAATTTGCGTCAAGGGAACTACTACGAATCCGCACTCACCGACTTTTGATCCCTTGGTTGACATGGATCCCAGACTTGTTGTTTCG
TTTCTCGATTTGCCTAGGGAAGCAGATGTGAGCTCGTTGGTCCTTAGGTTTGGTGGAGAATGTGAGCTGATATGGTTGAATGACAAGAATGCGTTGGCTG
TATTCAGTGACCCTGCTCGTGCAGCAACTGCTCTGAGGAGGTTGGATCATGCATCGGCATATTACGGAGTGGCTGTGGTTCTTCAGAACACTGGCCCGGG
TGGTGCCTGGGGAGCAATGGGTACCCCTGCTAGAGAAGGCGGAGGAGTGTCTGCCACATTGAAGGGGGCCAGTGCATGGAAGAAGGCTGTTACTCAGGTC
GACTCTTGGGGCGGAGAAGAACAACGACATGTAAGTTCCATGGATGCTCAAAGCACTCCCTGGAAACAAAAAGAAGCTGCCGTAAACGCTTCAGTCAATC
GTTGGAGTGTTCTGGACTTGGAATCGTCGTCCAGTAACGGTGCTTCGTTGATCAGAAGTAACAATCAAGGTGGAAGCCATTCTGGTTCGGGGGCTGGTGC
GAGTTTACTCCCTCGTGATGCCGTTGTCCCAGAGGGAGAGTCGTCAGAAGTGGTGGATGATTGGGAGAAAGCTTATGACATGGAGAATGATGTGCAGTAG
|
|||||||||||||||||||||||||
|
AA sequence
|
>Lus10025071 pacid=23167869 polypeptide=Lus10025071 locus=Lus10025071.g ID=Lus10025071.BGIv1.0 annot-version=v1.0
MSSQPRGDRRDRSRSSSQTAQQRWIPRGSAPSSVSVNNNIPPATFNPSNVDGRGEQSNANPNSGPREARPKGAGNASRSYVAGRPVNNHRRDKQREKGET
HAVKESVDPNLPQLVQEIREKLAKGTVECMICYDMVRRSAPIWSCSSCYSIFHLHCIKKWVRAPTSIDMSADKHQGFNWRCPGCQYVQLTPLKEISYLCF
CGKRTDPPSDLYLTPHSCGEPCGKPLERHLPGSLETKEDSCPHNCVLQCHPGPCPPCKSFAPPCLCPCGKKTITTRCSDRQSVLTCGQQCDKLLDCQRHR
CGRMCHVGPCDPCEILVTASCFCKKKQEVVYCGDMSVKGEVNVKDGVFSCNSSCGKLLACGNHECDEVCHPGPCGSCALMPGRVKSCYCGKTSLQKERKS
CLDPIPNCSQVCSKLLPCGIHNCEDLCHSGDCPPCMVSVSQKCQCGSTSRTVACYQTSTESEKFKCDKPCGRKKNCGRHRCSERCCPLSNPNPVRSEDWD
PHFCQMACGKKLRCGQHSCESLCHSGHCPPCLETIFSDLTCACGRTSIPPPLPCGTPAPSCQLPCSVPQPCGHLASHSCHYGDCPPCSVPVAKECVGGHV
VLKNIPCGSKDIRCNKLCGKTRQCGLHACGRTCHPAPCDSSGSTSAGAARVSCGQTCGAPRRDCRHTCAALCHPSSPCPDVRCQFPVTISCSCGRITASV
PCDAGGTNAGYSADTVFDASIAHKLPAPLQPVESTGSRIPLGQRKLTCDEECAKSERKRVLADAFDISTPNFEALHFGENSQVTELIADLYRRDPKWVLA
VEERCKYLVLGKSKGTVGGLKVHVFCPMMKDKRDAVRLIAERWKLSINVAGWEPKRFVVLHATPKSRPPSRVICVKGTTTNPHSPTFDPLVDMDPRLVVS
FLDLPREADVSSLVLRFGGECELIWLNDKNALAVFSDPARAATALRRLDHASAYYGVAVVLQNTGPGGAWGAMGTPAREGGGVSATLKGASAWKKAVTQV
DSWGGEEQRHVSSMDAQSTPWKQKEAAVNASVNRWSVLDLESSSSNGASLIRSNNQGGSHSGSGAGASLLPRDAVVPEGESSEVVDDWEKAYDMENDVQ
|
DESeq2's median of ratios [FLAX]
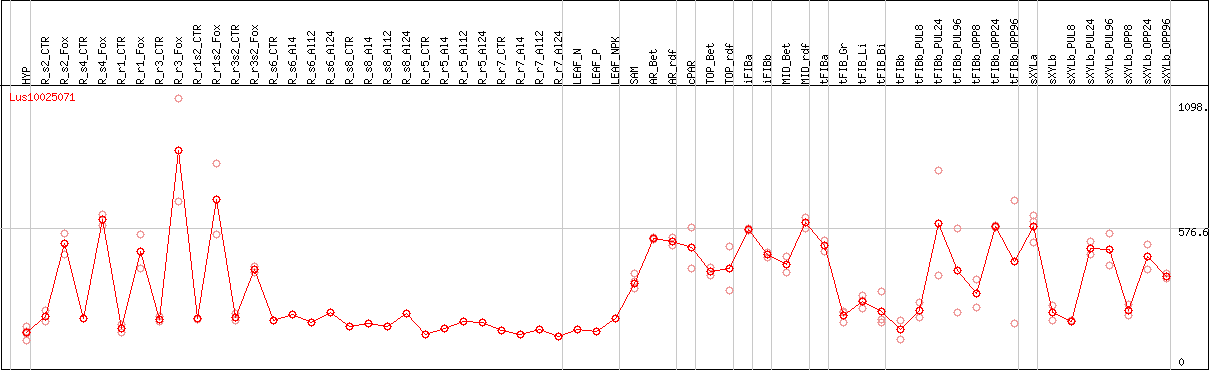
Coexpressed genes
Lus10025071 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
