External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13580 71 / 2e-15
ABCG6
ATP-binding cassette G6, ABC-2 type transporter family protein (.1)
AT2G39350 68 / 2e-14
ABCG1
ATP-binding cassette G1, ABC-2 type transporter family protein (.1)
AT3G53510 66 / 2e-13
ABCG20
ATP-binding cassette G20, ABC-2 type transporter family protein (.1)
AT2G37360 63 / 2e-12
ABCG2
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
AT3G55110 62 / 2e-12
ABCG18
ATP-binding cassette G18, ABC-2 type transporter family protein (.1)
AT3G55100 59 / 4e-11
ABCG17
ATP-binding cassette G17, ABC-2 type transporter family protein (.1)
AT1G31770 50 / 4e-08
ABCG14
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
AT1G51460 50 / 5e-08
ABCG13
ATP-binding cassette G13, ABC-2 type transporter family protein (.1)
AT1G51500 48 / 2e-07
AtABCG12, WBC12, ABCG12, D3, CER5, ATWBC12
ECERIFERUM 5, ARABIDOPSIS THALIANA WHITE-BROWN COMPLEX 12, ATP-binding cassette G12, ABC-2 type transporter family protein (.1)
AT3G52310 48 / 3e-07
ABCG27
ATP-binding cassette G27, ABC-2 type transporter family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030307
75 / 6e-17
AT2G37360 366 / 9e-122
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Lus10039859
57 / 2e-10
AT2G37360 319 / 5e-97
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Lus10018624
57 / 2e-10
AT2G37360 316 / 7e-96
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Lus10028704
51 / 3e-08
AT1G71960 818 / 0.0
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
Lus10004274
50 / 4e-08
AT2G37360 296 / 2e-90
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Lus10019238
50 / 4e-08
AT2G37360 296 / 1e-90
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Lus10034775
50 / 5e-08
AT1G31770 1028 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Lus10033315
50 / 5e-08
AT1G31770 551 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Lus10023154
50 / 6e-08
AT1G17840 613 / 0.0
DESPERADO, CUTICULAR DEFECT AND ORGAN FUSION 1, ARABIDOPSIS THALIANA WHITE-BROWN COMPLEX HOMOLOG PROTEIN 11, ATP-binding cassette G11, white-brown complex homolog protein 11 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G213400
78 / 6e-18
AT3G55090 1014 / 0.0
ATP-binding cassette G16, ABC-2 type transporter family protein (.1)
Potri.008G047900
74 / 1e-16
AT3G55090 1008 / 0.0
ATP-binding cassette G16, ABC-2 type transporter family protein (.1)
Potri.002G156900
65 / 2e-13
AT3G53510 950 / 0.0
ATP-binding cassette G20, ABC-2 type transporter family protein (.1)
Potri.014G080200
65 / 2e-13
AT2G37360 955 / 0.0
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Potri.006G215100
59 / 3e-11
AT3G53510 860 / 0.0
ATP-binding cassette G20, ABC-2 type transporter family protein (.1)
Potri.012G045100
56 / 3e-10
AT3G55130 270 / 3e-79
ATP-binding cassette G19, white-brown complex homolog 19 (.1)
Potri.015G036100
55 / 7e-10
AT2G37360 276 / 2e-81
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Potri.013G111900
54 / 2e-09
AT1G71960 795 / 0.0
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
Potri.019G083000
53 / 4e-09
AT1G71960 782 / 0.0
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
Potri.003G046800
53 / 4e-09
AT1G31770 945 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
PFAM info
Representative CDS sequence
>Lus10025149 pacid=23167751 polypeptide=Lus10025149 locus=Lus10025149.g ID=Lus10025149.BGIv1.0 annot-version=v1.0
ATGAGTTTTTTCACCGCGACGAAGACGCTTGTCAATGATATCTATGGAGAAGCAAGTGACGGTGAGATTTTGGTCGCTCTCGGACCGAGTGGCTCCGGGA
AATCAACTCTGCTCGATGCTCTGGCGAATCGAATTGCAAAAAAATTGTTGAAAGGGAGAGACATTTATGTACGCGACAGAGTTACGGCTCTCGAAGACTC
TGTCCACGTCGAAGAACAAGGCTCGAGTCCAGGCGCTGATCGACCAGCTCGGACTCCGGAAAGCCACAAAAATGGTCATCGGAGGCGAAGGCCACCGGGA
AGTCTCCGACAGTCGGTGGAGGCAAAGATACCGCCGATGCGGTAG
AA sequence
>Lus10025149 pacid=23167751 polypeptide=Lus10025149 locus=Lus10025149.g ID=Lus10025149.BGIv1.0 annot-version=v1.0
MSFFTATKTLVNDIYGEASDGEILVALGPSGSGKSTLLDALANRIAKKLLKGRDIYVRDRVTALEDSVHVEEQGSSPGADRPARTPESHKNGHRRRRPPG
SLRQSVEAKIPPMR
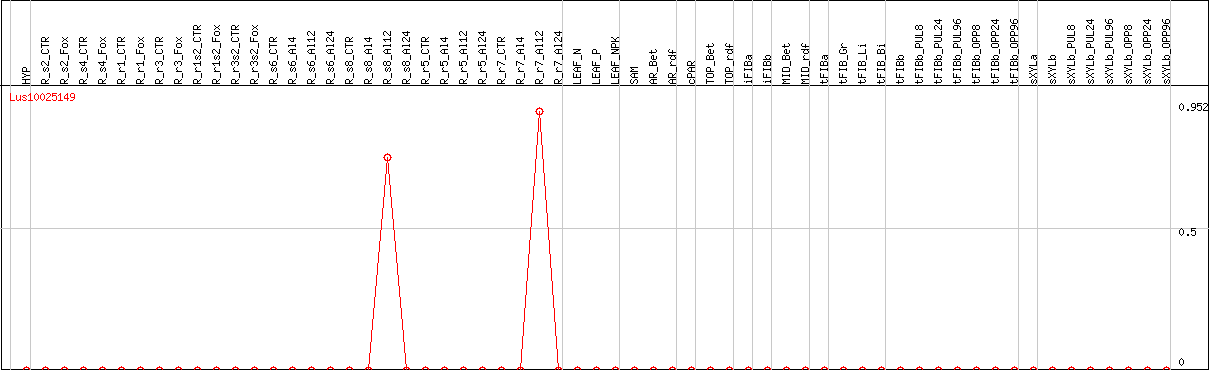
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025149 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.