External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G55090 52 / 2e-09
ABCG16
ATP-binding cassette G16, ABC-2 type transporter family protein (.1)
AT2G39350 52 / 2e-09
ABCG1
ATP-binding cassette G1, ABC-2 type transporter family protein (.1)
AT3G55110 40 / 3e-05
ABCG18
ATP-binding cassette G18, ABC-2 type transporter family protein (.1)
AT3G55130 39 / 0.0001
ABCG19, ATWBC19
ATP-binding cassette G19, white-brown complex homolog 19 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023465
135 / 2e-38
AT2G39350 1064 / 0.0
ATP-binding cassette G1, ABC-2 type transporter family protein (.1)
Lus10040341
130 / 4e-37
AT2G39350 452 / 0.0
ATP-binding cassette G1, ABC-2 type transporter family protein (.1)
Lus10001971
103 / 3e-30
AT3G55090 79 / 3e-18
ATP-binding cassette G16, ABC-2 type transporter family protein (.1)
Lus10030307
102 / 1e-27
AT2G37360 366 / 9e-122
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Lus10025281
44 / 3e-06
AT2G37360 1035 / 0.0
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G047900
96 / 1e-24
AT3G55090 1008 / 0.0
ATP-binding cassette G16, ABC-2 type transporter family protein (.1)
Potri.010G213400
86 / 4e-21
AT3G55090 1014 / 0.0
ATP-binding cassette G16, ABC-2 type transporter family protein (.1)
Potri.014G080200
49 / 4e-08
AT2G37360 955 / 0.0
ATP-binding cassette G2, ABC-2 type transporter family protein (.1)
Potri.006G215100
46 / 5e-07
AT3G53510 860 / 0.0
ATP-binding cassette G20, ABC-2 type transporter family protein (.1)
Potri.002G156900
43 / 6e-06
AT3G53510 950 / 0.0
ATP-binding cassette G20, ABC-2 type transporter family protein (.1)
PFAM info
Representative CDS sequence
>Lus10025150 pacid=23167774 polypeptide=Lus10025150 locus=Lus10025150.g ID=Lus10025150.BGIv1.0 annot-version=v1.0
ATGGAACTCACCGATCATCAGCCTCCGCCACCGACATCTTCCCGACATGGAGCCTTATGCCTTTCTCCAACACTCGGCCAGCTTCTGAAGCACGTTGGAG
ATGTTCTAAAGGAAGTCACCGGTGATGGAAGCGAAACTCCCGTCCATCACGTGCCCGAGCTCGGGGATTCTAATCCGAAAGTTTCCCGATCGATTCCGTT
CGTTCTGTCTTTCAGTAATCTTACTTACACGTTCGCATCCACCGCAAGCTGA
AA sequence
>Lus10025150 pacid=23167774 polypeptide=Lus10025150 locus=Lus10025150.g ID=Lus10025150.BGIv1.0 annot-version=v1.0
MELTDHQPPPPTSSRHGALCLSPTLGQLLKHVGDVLKEVTGDGSETPVHHVPELGDSNPKVSRSIPFVLSFSNLTYTFASTAS
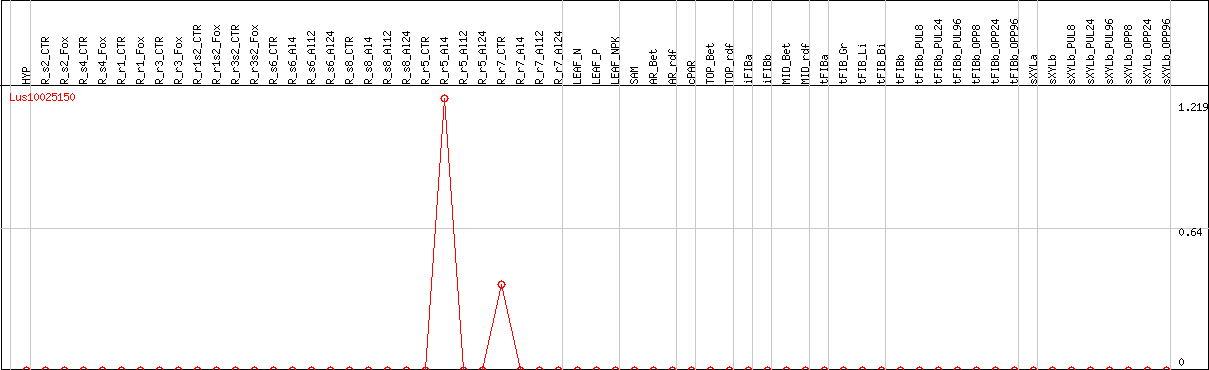
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025150 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.