Lus10025168 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10025168 pacid=23167747 polypeptide=Lus10025168 locus=Lus10025168.g ID=Lus10025168.BGIv1.0 annot-version=v1.0
ATGGGTCAGGCAAATGAGGGCCTGAATCTTAGTGTTCAGGATATTGATGCTTACTGGCTGCAGAGGAAGATCTCTCAGGCTTATGAACAACAGATCGACC
CACAGCAGTGTCAAAAGCTTGCTGAAGAGGTGTTAAAGATACTTGCTGAGGGAGATGATAGGGAAGTTGAGACGAAGTTGGTGGTGCTCCTTGACTTTGA
AAAGTTCAGCCTTATAAAATTTCTCTTACATAATAGACTGAAGATCGTGTGGTGTACCCGATTGACTCGTGCGAAGGATCAAGAAGAGAGGAACCATATT
GAAGAGGAAATGATGAACTCTGGTGCAGATTTGGCTGCGATTCTACAACAGCTTCGAGCTACACGAGCAACTGCAAAAGAAAGGCAAAAGAACCTGGAGA
AGAGCATAAGGGAAGAGGCTCGCAGACTCAAGGATGATATTGTCGGTGATGAGGGTAAAGACAGTACTGCTCTTGTGGACAGAGATGCAAACAGTGGGTG
GGTGAAGAGCCAACTCAAGCTGCTAAACCTGGACAATATTGCTTTCGAACAGGGCGGTCTTCTGATGGCAAATAAGAAGTGTGATCTTCCTGAAGGTTCT
TACAGACATAATCGCAAGGGTTTTGAAGAAGTTCACGTGCCAGCTTTGAAACCAAAACCTCTGGGTCCCGATGAGAAACTTGTGAGAATATCTGAGATGC
CAGAATGGGCACAACCTGCTTTTAAAGGTATGCAACAACTGAACAGGGTGCAGAGTAAGGTCTATCAGACTGCTTTCTTCTGTGCCGACAACATTCTCTT
GTGTGCGCCAACTGGGGCTGGGAAAACCAATGTTGCAGTTCTGACGATACTTCAGCAGCTTGGCCTTTATAGGAACAAGGATGGATCGTTTGATCATAAC
GGCTATAAAATAGTTTATGTGGCGCCCATGAAAGCTCTTGTGGCTGAAGTTGTTGGCAATTTGTCCAATCGTCTGCAAGAATACGGGGTGAATGTGAAGG
AACTAAGTGGAGATCAGACACTGACTCGTCAACAAATTGAAGAAACTCAGATAATTGTGACTACTCCAGAGAAATGGGATATCATTACCAGGAAATCTGG
GGATCGCACTTACACTCGTCTAGTAAAACTTCTTATAATTGATGAGATCCATCTTCTTCATGATAACAGAGGTCCTGTACTGGAAAGTATAGTTGCTAGA
ACTATTAGACAGGTTGAAACAACAAGGGACCATATCAGAATGGTGGGGTTGTCAGCTACTCTGCCGAACTACGAAGATGTGGCACTTTTCTTAAGAGTTG
ACGTGGGAAAAGGCCTTTTCCACTTTGATAATAGCTACAGACCTGTTCCTCTTGCACAACAGTATATTGGTGTCTCAGTCAAGAAGCCATTGCAGAGATT
TCAGTTAATGAATGATCTTTGTTACGAGAAGGTGATAGCTGTAGCTGGAAAGCACCAAACTCTCATTTTTGTTCACTCAAGAAAGGAAACAGCTAAAACT
GCGCGAGCAGTACGAGACACTGCACTTGGTAAGGATACTCTCGGAAGATTCTTGAAGGAGGACAGTGCTAGCCGTGAGATCCTCCAGAGTCATACAGATA
TGGTTAAAAGTAATGATCTGAAAGATCTTTTGCCGTATGGCTTTGCGGTTCATCATGCAGGTTTGTCAAGGCCTGATCGTCAAATTGTGGAGGATCTTTT
TGCCGGTGGGCATATACAAGTTTTGGTTTCTACTGCAACTCTAGCTTGGGGTGTAAATTTGCCTGCTCATACTGTGATCATTAAAGGGACTCAGATTTAC
AATCCAGAAAAAGGTGCATGGACTGAACTAAGTCCACTTGATGTTATGCAGATGTTAGGTCGTGCAGGTCGACCACAGTATGATACCTATGGTGAAGGTA
TTATCATCACGGGTCACAGTGAGCTGCAGTATTATCTTTCTTTGATGAATCAACAGCTTCCTATTGAAAGCCAATTCATATCCAAACTGGCTGATCAGTT
GAATGCAGAAATTGTTCTTGGTACTGTTCAGAATGCTAGAGAAGCCTGCAATTGGCTTGGTTATACATATTTGTATGTTCGAATGTTGAGGAATCCTACC
CTTTATGGTCTACCAGCTGATATTCTTACCAGAGATATTACACTGGAGGAGAGGAGGGCTGATTTGGTTCATTCAGCTGCTACAATTTTGGACAAGAATA
ACTTGGTCAAGTATGACCGGAAAAGTGGATATTTCCAGGTTACCGACCTTGGTCGGATTGCCAGCTACTATTATATTACACACGGCACCATATCCACGTA
TAATGAGCATCTGAAGCCCACTATGGGTGACATTGAGCTCTGTCGCTTGTTCTCACTCAGCGAAGAATTTAAGTATGTTACAGTGAGACAAGATGAGAAA
ATGGAACTAGCAAAGCTGTTGGATCGTGTTCCCATTCCAGTCAAGGAGAGTCTGGAGGAGCCTAGTGCCAAGATTAATGTTCTTCTTCAAGCTTACATAT
CACAGCTGAAGCTTGAAGGTCTGTCTTTGACTACTGACATGGTTTATATAACTCAGAGTGCGGGTCGTCTTCTTCGAGCACTTTTTGAGATTGTATTGAA
AAGAGGGTGGGCGCAATTGACTGAGAAAGCTTTAAACTTATGTAAGATGGTTAACAAGAAGATGTGGAGTGTTCAGACCCCCCTTCGTCAATTTAATGGG
ATACAGAAAGAGATTCTTATGAAATTGGAAAAGAAAGACTTGTCATGGGAAAGGTACTATGATCTTACATCCCAGGAAATTGGGGAACTAATCCGTCTCC
CAAAAATGGGCAGAGTACTGCATAAGTTCATCCATCAGTTCCCAAAGCTGAACCTTGCAGCTCATGTGCAGCCAATTACACGAACTATTTTGAGGGTAGA
GCTCACCATAACACCTGACTTCCAGTGGGATGACAGGGTTCATGGGTATGTGGAGCCATTTTGGGTTATTGTTGAGGACAACGATGGGGAGGATATCCTG
CACCACGAGTATTTTATGTTGAAGAAGCAGTATGTTGATGAGGACCACACGTTGAATTTCACAGTGCCAATCTATGAGCCGCTTCCACCTCAATACTTCA
TCCGTATTGTATCTGATAGGTGGCTTGGATCGCAAACTGTCCTGCCTGTATCCTTCCGGCACCTAATTTTACCTGAAAAGTATCCACCCCCAACTGAGCT
GTTGGATTTGCAGCCTCTTCCAGTGACAGCTTTGAGAAATCCTATATATGAAGCTCTGTATAAGGGTTTTAAGCATTTTAATCCAGTCCAGACTCAGGTG
TTCACTGTTCTGTATAATACAGATGACAATGTCTTGGTTGCTGCACCAACCGGGAGTGGGAAGACGATATGTGCTGAGTTTGCAATATTGAGGAACTACC
AAAAGGTACCCGATAGTGTTATGCGAGCCGTTTACATTGCTCCCCTTGAAGCAATTGCTAAAGAGAGGTATGGCGATTGGAAAGAGAAGTTTGGTAAAGG
GCTTGGGTTGACAGTCGTGGAACTCACTGGGGAAACGGCTACTGATCTGAAACTCATTGAGAAGGGCCAAATAATTGTAAGTACTCCAGAGAAATGGGAT
GCTTTGTCCCGTCGTTGGAAGCAGAGGAAGTATGTTCAACAAGTTAGTCTGTTCGTGACCGATGAGCTTCATTTGATAGGCTTTTTGCTTCATTTGATAG
GCGGTCAGGGTGGTCCCGTGCTGGAGGTGATTGTCTCTCGGATGAGATATATTGCTAGTCAGACTGAGAACAAGATAAGAATTGTGGCTCTTTCATCTTC
TCTTGCCAATGCAAAAGATCTTGGAGAATGGATAGGTGCTACCTCGCACGGTCTTTTCAATTTCCCACCTGGTGTTCGTCCAGTACCCCTTGAGATACAC
ATTCAAGGGGTGGATATTGCTAATTTCGAGGCGAGGATGCAAGCTATGACCAAGCCAACTTATACTGCAATAGTTCAGCATGCAATGAACGGGAAGAATC
GGAAGCCTGCTATTGTCTATGTTCCAACTAGGAGATATGTGAAAGTCACTGCAGAGGACTTGATGTCATACTCGGCAGTTGATGCTGGAGAGCAGCCTGC
CTTTTTGTTACAACCTGTTGAAGAACTCGAGCCTTTCATAAATAAAATTAACGATGAAACTTTGAAGGTCACTTTGACATTTGGGGTTGGCTTCTTACAT
GAAGGCTTGAGCAGTTCAGATCAGGAGGTTGTTTCCCGGTTGTTTGAAGTTGGGTGGATTCAGGTCTGTGTTGTGAGCAGTTCAATGTCATGGGGAGTGC
CTTTGTTAGCACACTTGGTGGTTGTGATGGGAACTCAGTACTACGATGGCAGGGAAAATGCTCACACTGACTATCCAGTTACCGATCTTTTACAGATGAT
GGGCCATGCCAGTCGACCTTTGCTGGATAATTCCGGAAAGTGCGTCATTCTTTGTCACGCACCTCGCAAAGAATACTATAAGAAGTTTCTATATGAGGCC
TTCCCTGTGGAAAGTCACTTGCACCATTTTCTGCATGATAACTTCAATGCTGAGATAGTTGCTGGAGTTATTGAGAACAAGCAGGATGCTGTAGACTATC
TTACATGGACTCTCATGTATAGGAGGTTGACTCAGAATCCAAATTACTACAATCTTCAAGGTGTGAGTCACAGGCATCTTTCTGACCATCTCTCAGAACT
TGTGGAGAATACCTTGAGCGATTTGGAAGCAAGTAGGTGCATTTCAATCGAGGATGACATGGACCTTTCTCCCCTGAATCTCGGCATGATAGCATCTTAC
TATTACATCAGCTATACGACGATTGAGCGTTTCAGCTCGTCCTTGACTCCTAAAACAAAGATGAGGGGTCTTCTCGAGATTCTGGCTACTGCCTCAGAGT
ATGGCCAGCTTCCAGTAAGACCTGGAGAGGAGGAGAAACTCCGAAGGCTGTTGAACCACCAGAGATTCTCTATTGAAAGTCCAAGATACAGCGACCCCCA
GGTGAAGGCCAATGTTCTGCTCCAAGCTCATTTTTCGAGGCAAACAGTTGGCAAGAATCTGGCTCTTGACCAGCGAGAGGTTCTCCTCTCTGCTGGTCGA
TTACTTCAAGCAATGGTAGATGTCATATCGAGCAACGGTTGGTTAAACCTTGCTCTTCTCGCCATGGAAGTCAGCCAGATGGTAACACAAGGGATGTGGG
AGCGAGATTCCGTACTGATGCAGCTTCCACACTTTACAAGGGAGCTGGCCAAGAAGTGTCAGGAGAATCCCGGGAAGAACATAGAGACTGTTTTCGATTT
GGTGGAGATGGAGGATGAAGAGAGGCGCGAGCTTCTGCAGATGTCTGACCCCGAGTTGCTGGACATAGCAAAGTTCTGTAATCGCTACCCGAACATCGAT
ATGGTTTACGAGGTGGTGGATGCTGAGAATGTGAGGGCAGGGGATGACGTGACCTTACAGGTGGTGCTGGAGAGGGATCTTGAAGGAAGGACGGAGGTGG
GACCCGTGGAGACATTGAGGTACCCGAAGGCTAAGGAAGAAGGATGGTGGCTGGTGGTCGGTGATACCAAGACGAATCAGCTGCTGGCGATCAAGAGGGT
GTCACTGCAGAGGAAGCTGAAAGTGAAGCTGGAATTTGCGGCGCCTGCTGAAGCTGGAAAGAAAAGCTACACGCTCTACTTCATGTGCGATTCTTATCTG
GGTTGCGACCAAGAGTATGGTTTTAGTGTTGATGTTGGTGCACCTGAGGAGGAGGATGGTGCAAGAGAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10025168 pacid=23167747 polypeptide=Lus10025168 locus=Lus10025168.g ID=Lus10025168.BGIv1.0 annot-version=v1.0
MGQANEGLNLSVQDIDAYWLQRKISQAYEQQIDPQQCQKLAEEVLKILAEGDDREVETKLVVLLDFEKFSLIKFLLHNRLKIVWCTRLTRAKDQEERNHI
EEEMMNSGADLAAILQQLRATRATAKERQKNLEKSIREEARRLKDDIVGDEGKDSTALVDRDANSGWVKSQLKLLNLDNIAFEQGGLLMANKKCDLPEGS
YRHNRKGFEEVHVPALKPKPLGPDEKLVRISEMPEWAQPAFKGMQQLNRVQSKVYQTAFFCADNILLCAPTGAGKTNVAVLTILQQLGLYRNKDGSFDHN
GYKIVYVAPMKALVAEVVGNLSNRLQEYGVNVKELSGDQTLTRQQIEETQIIVTTPEKWDIITRKSGDRTYTRLVKLLIIDEIHLLHDNRGPVLESIVAR
TIRQVETTRDHIRMVGLSATLPNYEDVALFLRVDVGKGLFHFDNSYRPVPLAQQYIGVSVKKPLQRFQLMNDLCYEKVIAVAGKHQTLIFVHSRKETAKT
ARAVRDTALGKDTLGRFLKEDSASREILQSHTDMVKSNDLKDLLPYGFAVHHAGLSRPDRQIVEDLFAGGHIQVLVSTATLAWGVNLPAHTVIIKGTQIY
NPEKGAWTELSPLDVMQMLGRAGRPQYDTYGEGIIITGHSELQYYLSLMNQQLPIESQFISKLADQLNAEIVLGTVQNAREACNWLGYTYLYVRMLRNPT
LYGLPADILTRDITLEERRADLVHSAATILDKNNLVKYDRKSGYFQVTDLGRIASYYYITHGTISTYNEHLKPTMGDIELCRLFSLSEEFKYVTVRQDEK
MELAKLLDRVPIPVKESLEEPSAKINVLLQAYISQLKLEGLSLTTDMVYITQSAGRLLRALFEIVLKRGWAQLTEKALNLCKMVNKKMWSVQTPLRQFNG
IQKEILMKLEKKDLSWERYYDLTSQEIGELIRLPKMGRVLHKFIHQFPKLNLAAHVQPITRTILRVELTITPDFQWDDRVHGYVEPFWVIVEDNDGEDIL
HHEYFMLKKQYVDEDHTLNFTVPIYEPLPPQYFIRIVSDRWLGSQTVLPVSFRHLILPEKYPPPTELLDLQPLPVTALRNPIYEALYKGFKHFNPVQTQV
FTVLYNTDDNVLVAAPTGSGKTICAEFAILRNYQKVPDSVMRAVYIAPLEAIAKERYGDWKEKFGKGLGLTVVELTGETATDLKLIEKGQIIVSTPEKWD
ALSRRWKQRKYVQQVSLFVTDELHLIGFLLHLIGGQGGPVLEVIVSRMRYIASQTENKIRIVALSSSLANAKDLGEWIGATSHGLFNFPPGVRPVPLEIH
IQGVDIANFEARMQAMTKPTYTAIVQHAMNGKNRKPAIVYVPTRRYVKVTAEDLMSYSAVDAGEQPAFLLQPVEELEPFINKINDETLKVTLTFGVGFLH
EGLSSSDQEVVSRLFEVGWIQVCVVSSSMSWGVPLLAHLVVVMGTQYYDGRENAHTDYPVTDLLQMMGHASRPLLDNSGKCVILCHAPRKEYYKKFLYEA
FPVESHLHHFLHDNFNAEIVAGVIENKQDAVDYLTWTLMYRRLTQNPNYYNLQGVSHRHLSDHLSELVENTLSDLEASRCISIEDDMDLSPLNLGMIASY
YYISYTTIERFSSSLTPKTKMRGLLEILATASEYGQLPVRPGEEEKLRRLLNHQRFSIESPRYSDPQVKANVLLQAHFSRQTVGKNLALDQREVLLSAGR
LLQAMVDVISSNGWLNLALLAMEVSQMVTQGMWERDSVLMQLPHFTRELAKKCQENPGKNIETVFDLVEMEDEERRELLQMSDPELLDIAKFCNRYPNID
MVYEVVDAENVRAGDDVTLQVVLERDLEGRTEVGPVETLRYPKAKEEGWWLVVGDTKTNQLLAIKRVSLQRKLKVKLEFAAPAEAGKKSYTLYFMCDSYL
GCDQEYGFSVDVGAPEEEDGARE
|
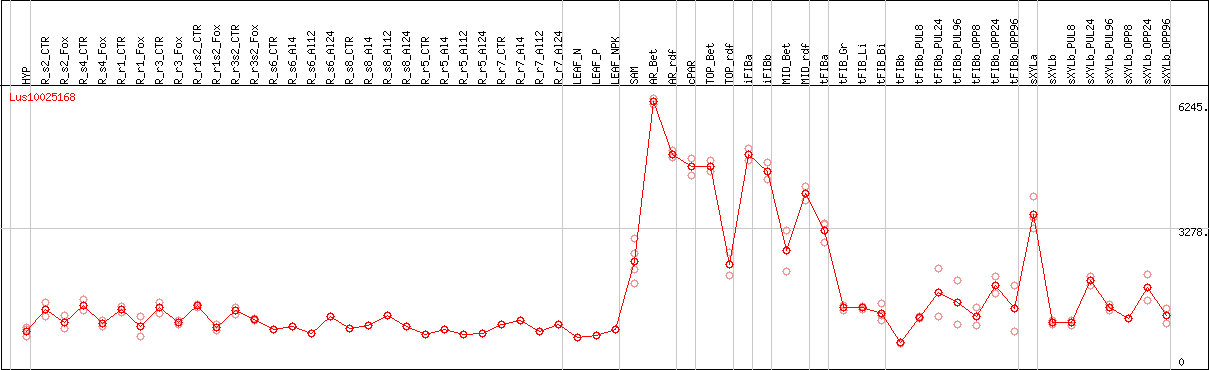
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025168 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
