External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G59320 99 / 5e-28
LTP3
lipid transfer protein 3 (.1)
AT5G59310 97 / 3e-27
LTP4
lipid transfer protein 4 (.1)
AT2G38540 91 / 8e-25
ATLTP1, LP1
ARABIDOPSIS THALIANA LIPID TRANSFER PROTEIN 1, lipid transfer protein 1 (.1)
AT3G51590 90 / 4e-24
LTP12
lipid transfer protein 12 (.1)
AT3G51600 87 / 5e-23
LTP5
lipid transfer protein 5 (.1)
AT3G08770 84 / 4e-22
LTP6
lipid transfer protein 6 (.1.2)
AT4G33355 84 / 6e-22
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
AT2G38530 83 / 1e-21
cdf3, LP2, LTP2
cell growth defect factor-3, lipid transfer protein 2 (.1)
AT5G01870 82 / 3e-21
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT2G15050 80 / 2e-20
LTP7, LTP
lipid transfer protein 7, lipid transfer protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025151
199 / 1e-67
AT5G59320 103 / 1e-29
lipid transfer protein 3 (.1)
Lus10022745
116 / 2e-34
AT5G59310 103 / 1e-29
lipid transfer protein 4 (.1)
Lus10014167
115 / 3e-34
AT5G59310 102 / 2e-29
lipid transfer protein 4 (.1)
Lus10025234
100 / 4e-28
AT2G38530 118 / 2e-35
cell growth defect factor-3, lipid transfer protein 2 (.1)
Lus10015279
89 / 6e-24
AT5G59320 116 / 6e-35
lipid transfer protein 3 (.1)
Lus10015278
88 / 1e-23
AT5G59310 110 / 1e-32
lipid transfer protein 4 (.1)
Lus10026418
87 / 7e-23
AT5G59320 115 / 2e-34
lipid transfer protein 3 (.1)
Lus10025148
87 / 7e-23
AT5G01870 105 / 6e-30
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10000082
85 / 3e-22
AT2G38540 88 / 2e-23
ARABIDOPSIS THALIANA LIPID TRANSFER PROTEIN 1, lipid transfer protein 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G086500
103 / 1e-29
AT2G38540 124 / 5e-38
ARABIDOPSIS THALIANA LIPID TRANSFER PROTEIN 1, lipid transfer protein 1 (.1)
Potri.004G086600
102 / 4e-29
AT2G38540 122 / 3e-37
ARABIDOPSIS THALIANA LIPID TRANSFER PROTEIN 1, lipid transfer protein 1 (.1)
Potri.016G135400
84 / 6e-22
AT5G59320 109 / 3e-32
lipid transfer protein 3 (.1)
Potri.016G136000
84 / 1e-21
AT5G01870 113 / 1e-33
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.006G108100
83 / 2e-21
AT2G38540 121 / 6e-37
ARABIDOPSIS THALIANA LIPID TRANSFER PROTEIN 1, lipid transfer protein 1 (.1)
Potri.016G135800
82 / 3e-21
AT5G01870 111 / 1e-32
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.001G232900
81 / 6e-21
AT2G18370 100 / 3e-28
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.001G232700
81 / 1e-20
AT2G18370 93 / 1e-25
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.016G135500
78 / 1e-19
AT5G59320 94 / 8e-26
lipid transfer protein 3 (.1)
Potri.014G046500
75 / 2e-18
AT4G33355 76 / 6e-19
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0482
Prolamin
PF00234
Tryp_alpha_amyl
Protease inhibitor/seed storage/LTP family
Representative CDS sequence
>Lus10025231 pacid=23157363 polypeptide=Lus10025231 locus=Lus10025231.g ID=Lus10025231.BGIv1.0 annot-version=v1.0
ATGGAGGCTAGTCATATATTTGTTACAATGCTCCTCGTCGTCGGCGCACTGACAGTGGCTACTGGAGCGGACGACGGAGGAATAACGTGCGGTGAAGTGG
CGATGAGCATGAGGCCATGCGTACCTTACTTGCTAAAAGGTGGTCCCGTGCCGGCACCGTGCTGCAGCGGGATGAGGTCACTGAACGAGGCAGCCAAGAC
AACCGTTGATCAACAAAAGACTTGCGAGTGCTTGAAATCGGCGGCGGATAATATCCCTGACTTGAAGAACGAACTCGCTGCCGCACTCCCAGATGCCTGT
GGGGTCGACTTTCCGTACGAGTTCACCACCAAGACCGACTGCAATTCTGTGTTTTGTTCTGATCGTCGTGGCAGCATCGAGTAA
AA sequence
>Lus10025231 pacid=23157363 polypeptide=Lus10025231 locus=Lus10025231.g ID=Lus10025231.BGIv1.0 annot-version=v1.0
MEASHIFVTMLLVVGALTVATGADDGGITCGEVAMSMRPCVPYLLKGGPVPAPCCSGMRSLNEAAKTTVDQQKTCECLKSAADNIPDLKNELAAALPDAC
GVDFPYEFTTKTDCNSVFCSDRRGSIE
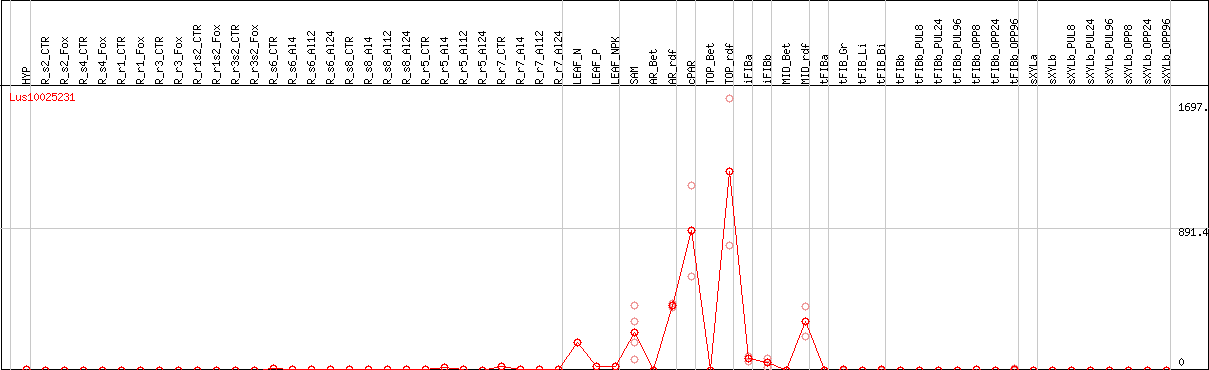
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025231 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.