External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G01470 54 / 1e-09
HD
HD-ZIP-1, HAT5, ATHB1, ATHB-1
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
AT1G69780 47 / 4e-07
HD
ATHB13
Homeobox-leucine zipper protein family (.1)
AT2G22430 46 / 6e-07
HD
ATHB6
homeobox protein 6 (.1)
AT5G65310 44 / 2e-06
HD
ATHB-5, ATHB5
homeobox protein 5 (.1.2)
AT4G40060 44 / 3e-06
HD
ATHB16 ,ATHB-16
homeobox protein 16 (.1)
AT3G01220 44 / 5e-06
HD
ATHB20
homeobox protein 20 (.1)
AT1G27045 43 / 8e-06
ATHB54
homeobox protein 54, Homeobox-leucine zipper protein family (.1)
AT5G15150 43 / 9e-06
HD
ATHB3, HAT7, ATHB-3
HOMEOBOX FROM ARABIDOPSIS THALIANA 7, homeobox 3 (.1)
AT5G03790 43 / 9e-06
HD
LMI1, ATHB51
LATE MERISTEM IDENTITY1, homeobox 51 (.1)
AT2G36610 39 / 0.0002
HD
ATHB22
ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 22, homeobox protein 22 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036709
147 / 5e-45
AT3G01470 139 / 2e-39
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10037218
145 / 2e-44
AT3G01470 140 / 7e-40
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10013277
91 / 3e-23
AT3G01470 130 / 2e-36
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10030800
91 / 3e-23
AT3G01470 135 / 4e-38
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10008795
53 / 3e-09
AT3G01470 261 / 3e-87
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10022225
53 / 3e-09
AT3G01470 270 / 2e-90
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10004270
53 / 4e-09
AT3G01470 152 / 5e-44
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10042180
52 / 5e-09
AT3G01470 159 / 6e-47
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
Lus10036674
50 / 5e-08
AT3G01470 153 / 8e-45
HOMEODOMAIN PROTEIN FROM ARABIDOIPSIS THALIANA 5, ARABIDOPSIS THALIANA HOMEOBOX 1, homeobox 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00046
Homeodomain
Homeodomain
Representative CDS sequence
>Lus10025232 pacid=23157384 polypeptide=Lus10025232 locus=Lus10025232.g ID=Lus10025232.BGIv1.0 annot-version=v1.0
ATGGCGAGCGAAAACTCTTCGGCGTTAGCGGCGGCTTCGACCACCTCCCTTACTGGATCCCATCCATTTCCACCACTCTCTACGGTGAAATTTCAGACGG
CGGCGAGGCAGATGGTGGAATACGAAAATGACAGGAATTTCTTCAATCCCATGGTGAAAGAAGAGCTCTACTGCGACGAGGATTTCGAAAACGGTTCAAG
CCCTCGCTGCAAGAAGAGGCGGCTGACAGCGAGTCAGGTTCAGTTCCTGGATCGGAGCTTCAAGAACGAGAACAAGCTCGGGCCGGAGAGGAAGATCCAG
CTCGCTAACAAGATCGGATTGTAG
AA sequence
>Lus10025232 pacid=23157384 polypeptide=Lus10025232 locus=Lus10025232.g ID=Lus10025232.BGIv1.0 annot-version=v1.0
MASENSSALAAASTTSLTGSHPFPPLSTVKFQTAARQMVEYENDRNFFNPMVKEELYCDEDFENGSSPRCKKRRLTASQVQFLDRSFKNENKLGPERKIQ
LANKIGL
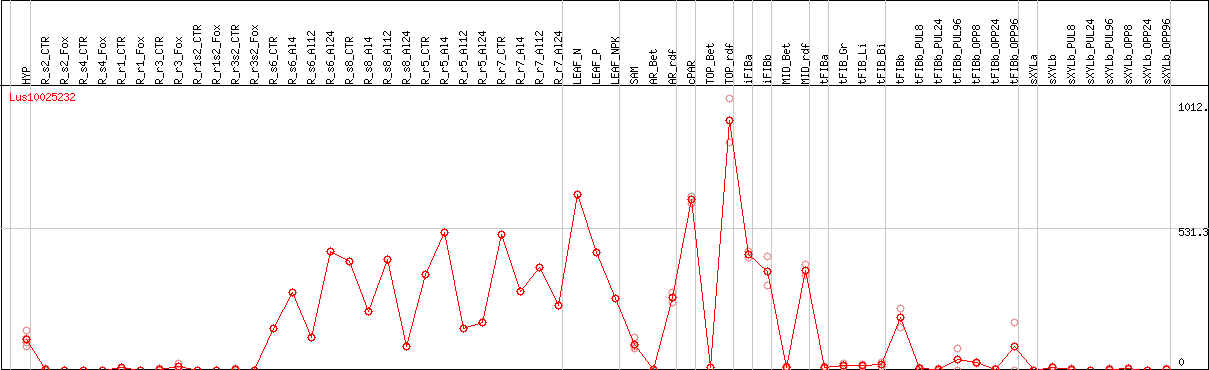
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025232 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.