External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G29390 194 / 4e-62
COR413IM2, COR314-TM2
COLD REGULATED 314 INNER MEMBRANE 2, cold regulated 314 thylakoid membrane 2 (.1.2)
AT1G29395 191 / 3e-61
COR413IM1, COR413-TM1, COR414-TM1
cold regulated 414 thylakoid membrane 1, COLD REGULATED 314 THYLAKOID MEMBRANE 1, COLD REGULATED 314 INNER MEMBRANE 1 (.1)
AT3G50830 60 / 9e-11
ATCOR413-PM2, COR413-PM2
cold-regulated 413-plasma membrane 2 (.1)
AT4G37220 59 / 2e-10
Cold acclimation protein WCOR413 family (.1)
AT2G23680 48 / 1e-06
Cold acclimation protein WCOR413 family (.1.2)
AT2G15970 42 / 0.0001
ATCYP19, FL3-5A3, ATCOR413-PM1, WCOR413-LIKE, COR413-PM1
CYCLOPHILLIN 19, ARABIDOPSIS THALIANA COLD-REGULATED413 PLASMA MEMBRANE 1, cold regulated 413 plasma membrane 1 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015258
392 / 2e-140
AT1G29390 223 / 1e-73
COLD REGULATED 314 INNER MEMBRANE 2, cold regulated 314 thylakoid membrane 2 (.1.2)
Lus10004599
263 / 5e-89
AT1G29395 201 / 9e-65
cold regulated 414 thylakoid membrane 1, COLD REGULATED 314 THYLAKOID MEMBRANE 1, COLD REGULATED 314 INNER MEMBRANE 1 (.1)
Lus10004547
240 / 4e-80
AT1G29395 140 / 3e-41
cold regulated 414 thylakoid membrane 1, COLD REGULATED 314 THYLAKOID MEMBRANE 1, COLD REGULATED 314 INNER MEMBRANE 1 (.1)
Lus10021020
63 / 5e-12
AT4G37220 258 / 3e-88
Cold acclimation protein WCOR413 family (.1)
Lus10027946
58 / 3e-10
AT3G50830 247 / 5e-84
cold-regulated 413-plasma membrane 2 (.1)
Lus10012034
57 / 7e-10
AT3G50830 250 / 4e-85
cold-regulated 413-plasma membrane 2 (.1)
Lus10027945
57 / 8e-10
AT4G37220 245 / 4e-83
Cold acclimation protein WCOR413 family (.1)
Lus10012035
56 / 3e-09
AT4G37220 242 / 6e-82
Cold acclimation protein WCOR413 family (.1)
Lus10016782
52 / 6e-08
AT3G50830 277 / 9e-96
cold-regulated 413-plasma membrane 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G353500
191 / 3e-61
AT1G29395 185 / 1e-58
cold regulated 414 thylakoid membrane 1, COLD REGULATED 314 THYLAKOID MEMBRANE 1, COLD REGULATED 314 INNER MEMBRANE 1 (.1)
Potri.007G033801
64 / 2e-12
AT3G50830 282 / 1e-97
cold-regulated 413-plasma membrane 2 (.1)
Potri.004G149100
59 / 2e-10
AT3G50830 250 / 4e-85
cold-regulated 413-plasma membrane 2 (.1)
Potri.009G109800
57 / 9e-10
AT3G50830 238 / 4e-80
cold-regulated 413-plasma membrane 2 (.1)
Potri.007G032000
41 / 0.0002
AT4G37220 72 / 7e-16
Cold acclimation protein WCOR413 family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05562
WCOR413
Cold acclimation protein WCOR413
Representative CDS sequence
>Lus10025378 pacid=23150815 polypeptide=Lus10025378 locus=Lus10025378.g ID=Lus10025378.BGIv1.0 annot-version=v1.0
ATGAGTATCGGCATCTCTTCGAAACCGCTGAGCTTTTGCCTCCAGAACCGCGTCAGTCTCACCGAGAACAAACAGAAACAACGGCTTCTCGCTCTCGACG
TCCGACGTTTCCGCAGCTCGGTTACTCCTCTTCGGGTTTCCAGCTCGCTCAGTTACAATCCTCTCAGTTTCGGAATCAAGAGCAACGACGATGCAATTTT
GAGGAAGAAGATAGTGGGAGGGAGGATGATCAGTACCGTCTGTTCTGCTGTGCCAGTCAGTTCTGCCCGGAACCTACAGTGGATCTCCACTATTGCTTCC
ATGCTGTTGATGCTTGCCAAAGGTACTGCTATCAACAAATGCTTTCTCGTTCCCTTCCTTGCATTGCAGGCACCAACTGCTGTGATATCTTGGGTTAAGG
GTGAATATGGTGTGTGGGCTGCATTTATGGCACTTCTAGTACGCCTCTTCTTCTTCATTCCTGGTGAACTAGAGTTGCCATTTATGGCTTTGCTTCTGAT
CATCTTAGCGCCGTATCAGGTTCTGAACTTAAAAGGGACACAGGAAGGTGCCATCATTAGCATAGCGATTGCAGCTTACCTGGCTTTCCAGCATTTTTCA
CGAGGAGGCAACATGCAGAGATCATTTGAGCAAGGTTCAATTGTTGCCACCGTAGCTATCATCTGTATTACAGTTGCCTCATTCTTGCTCTTGCTCTAA
AA sequence
>Lus10025378 pacid=23150815 polypeptide=Lus10025378 locus=Lus10025378.g ID=Lus10025378.BGIv1.0 annot-version=v1.0
MSIGISSKPLSFCLQNRVSLTENKQKQRLLALDVRRFRSSVTPLRVSSSLSYNPLSFGIKSNDDAILRKKIVGGRMISTVCSAVPVSSARNLQWISTIAS
MLLMLAKGTAINKCFLVPFLALQAPTAVISWVKGEYGVWAAFMALLVRLFFFIPGELELPFMALLLIILAPYQVLNLKGTQEGAIISIAIAAYLAFQHFS
RGGNMQRSFEQGSIVATVAIICITVASFLLLL
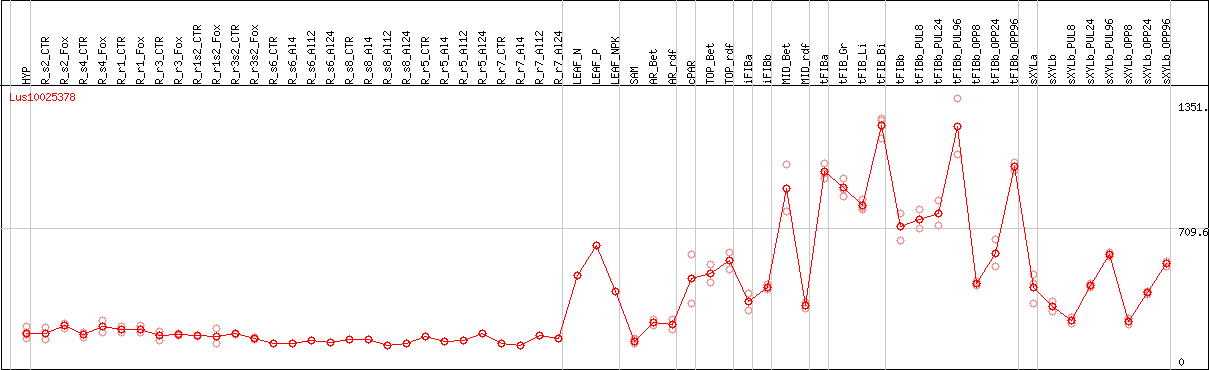
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025378 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.