Lus10025655 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10025655 pacid=23145021 polypeptide=Lus10025655 locus=Lus10025655.g ID=Lus10025655.BGIv1.0 annot-version=v1.0
ATGGACGCTACTCCTCTGAAACGTGTCTCCGAAGTAGGCGCCGAGCTCAGCCGTCTCACTCGCCCCGGCAAGGATACTCTCATCAAGTTACTCCGAGATG
GGGCGGATGCCTTAGCTCGAATTGAGCACACTTCGTTCCTCGAGAATTCGGCTAAAGCCGAAGCTCATAAGAAGCAAGCAACTGCGCTGAAACCACTGAA
GGAATCCATAGTTAAGCGCGGTCTTCAGCGCCATACTGACAGAGATGTCAAGCTCTTGGTGTCCGTCTGTGTCGCTGAGCTGTTCCGCATTCAAGCGCCG
GAGACTCCGTTTGCTGATAAGTATCTCGGAGATGTTTTCAAACTGCTGCTCGGTACGTTCTCGGAGCTTGCTGACGTTGATAGCCCTTACTTTGGGAAGA
GGGCCGGCGTTTTGGAGGCTGTAGCTAGATGCGAATGCTGTGTGATAATGCTGGATATCGGTTGTGGAGATTTGGTTGTCGAGATGTTCAATATTTTTTT
CTCAGTTGCGAGGGGACATCGTGGTAGTAGCGTGGAGAGGAATATCTTGCATATAATGTCTCGCATCTTAGACGAGGAGCCTTCTCAAAGACTTGTAGAT
ATAATTCTACAGAATCTAATAAAGGAAGAAGAGGGTCCGTCACGGCTTGCTGTTTCAATTATCAACAAGTTTCATGATACACTTAAGCCTTCCATTTGTC
GCTTCTTGAGTTCTTGCTTCTTGGAAAGAGATGCAGTTGAAAGTGAAGTAAAAGAATCTTACCATGATATATTATTTCAAGTTTACAAACACGAGCCTGA
GATGCTTCTTAATGTCATCCCTTACCTGACTGAAGAACTACGGACTGACCAGGTTGATGTCCGATTAAAAGCTGTCACTTTGCTTGGGAAACTACTTGCT
CTACGTGGGGACCATTTTGCTCAGCAACACCACAGCGTTCTCATGGAGTTCAAGAACAGGTTTTCTGATAAATCTGTGGGAGTTAGACTCAGTGCCCTTC
AATGTGCCAAAGCCAGTTACATTGCCAATCCACGGGGAAAATTATCAGAAGAGATTCTTGGCAAGTCCTTTTCCGCATTTGAGGGACGATTGTTGGACTT
CAGTGACGATCTCTCTGTTAGAAAGAAGGCCTTGGTGAAGTTGATGGAATTATATCGAGATTATTGCAGCAAATGTTCCGAACGGCGTATGGCTATTGAA
GATCACTTTGAACAGATTCCTCCTAAAGTTTTGATGCTATGGCATATAGACAGTAAGGAGTTCAGGTTCCAAATTGTGGAGCGTGTTCTTGGCGAGGACT
TATTTCCAGCTAATCTTTCTGCGGAAGAGAGGAGCAAACATTGGGTTCATATAGTTTCTCTTTTCACTCCTCCACACTGGAAGGTCCTGTCCTCTGTTCT
GTCTCAGAAAAACAGATTACAGACAGATATGCGAACTTATTTATCTTTGAGAATGAAAGCAAAGGGGACTGGTTTGGAAGTGATGGAGGGTAAAATAAAA
AAGTTAGTCATAAATATGTCAGCGTTGTTTCCTGATCCCTCCAAGGCTGCAGAATGCTTTAATAGTTTGAACCAAATGAAAGACTGCAATATTTTCAGTA
CCCTGGAGTTACTACTAAAAGATTCGATGATTACTGACGTGCAAGTGAATAGGGATAAATTTCAGAAAGTGATTGGAGACAAACATCCACAACATGATTT
TCTACAATTACTTTCTTCTAAGTGTTCTTTTACCATATTCAACTCAGAGATAATACGCTGTATCTTGGTTAATATTTCCAGCAACGTGCTTAACAGCAAG
CAAAAGGAGGCTTCTGCTAAACTCCTTATGGCCGTCATCAGTGCTTTCCCCTTTCTTTTAAGGGGTTCAGAAAAGGAAGTTCAGATGCTGTTGGGCGATA
ATACTCTCACAGCAGTGCTGATTGAAGTACTTGCAAAAGCGGGTTCCCACTTATCTGCCAACTTGAGTGATTTCTACCCGTTTCTTAGGAAAGTGTCTTT
AGAAGGAACTCGTGTTCAGGCGAAGCATGCCGTTTCAGCATTTGTTTCACTGGCTGGTTCTTCCGAGCAAGCTGCTCTATCACAACTTTGTGAGGAACTT
GTCAACTCTCTTCGTTGTGGGTGGAACACTTCAACTGTGTTACAGTCTCTGGGCTGTATTATGCTGCATTCTGTATCAGCATCTGAGATACATGGTGAAG
AGATCAGATCGTATATATTTGATATAATTTTCAAATTGAATCAATCTGACGATCCAATGTGTTCTGGGGAGATGTTGAACTGCACTGATTCCTGTAAGCT
GAAGATTTTTGCACTGAAAACACTTGTCAGGAGCGTTCTACCAGGGCAAGGATCTCTTTTAAAATGGAAAATTAATGAAATGCTGGGTCTTTTGTTGAAG
ATGCTACGGACAGGTGACACTTCTGATGGAGTCGTATCATGTGCAAGTGACAAGCCTTATATCCGATTAGCTGCTGCAAAATCTATTCTCCAGCTCTCGA
AGAGATGGGATTTACATATTTCTCATAAGATATTTCAGTTGACAATATTAATGGCAAAGGACCCTTCTCCTGTAGTTAGAAGATTATTCCTTTACAAAAC
GCACAAGCTATTGACGAATCATGCTATACCTAGTAGGTATGCATGTGCATTTGCATTGGCTGCATGTGGTCCCTGTGAAGATCTGCGTGAAGCTTCGTTT
AAATACATGAGAGACTTTATCAAATATTACGTAAGAATATCCCAATTTAAGCAAACGTCAGTGGGGCAAAGAGAATCCCCTACAGATGACCCTGCATTTA
TGGTTTTTTTCTTGATCCACATTCTTGCTCATGATGACGGCTTCCCACATGAGGATTGCCAAGATGAGCAAGTTTATGCTCGGTTCTGTAGTCCACTCTT
TTGGGCTATACAGGAATTAACAAGGTCAACTAATGCAAACGGCAGCTTGGATTTTGCTGATTATACCATCCTACGTTTGCTCTCTGTCTTCCGCGCAATC
AAAAGAGCAGAAGATGCTGTTGATCCTCAAGGCACTCCCAAACTGCAAATTCTAGCAGATATTGGGATCTCCATTGTGAACTCATTGCGGCATAAAGGCA
AGTCTGTATCACAGACGCCAGGACCAGTTTTGCTGCCATCATCATTATATAAATTAAGTCCAGTTAACTTGAAACGCATCAGCAGCTCCGCCGATCGTCA
AAATTTCTTCAGCCGGCTTATTCCAATTCTGAAGTCCGAAGTAGTCATCCCTGCCGAGACTTCTAATAAACGTGGTAGAGATTGTCATGAGGATAAGCTG
CATTCTGGAGATGTTGAGTTTAGTACTTCACTAAGTTCGGAATTGCGCGAGCCTATCCATTTGCAAAATCTGATTTCTACAGAAGTCCAGGCAACAGTGG
CTGGGGATTTCAGCTTGGGACGTTTATTAGAAGAAGCTCTTTCACCCACCTCTGAATCTGACCAATTATACAGTGATGACACTTCAACAGAGGTCAACCA
AGGTCGTGCCTGTAAGTTTCAAAAGCCCACATTCTCCAGTTCGCCCATATTGTCTGGTTCTGCAATTGCCACGCCTTCTGTGCTGCAGTGTGATGTACTA
CAGCAAAAGTCGGGCGGAAATGTTTCATCGTTGCAAGTGACTAGCGGTACTGCTGATAATAGTGCGGAGATGGAGTTGATACGTTTGGATGATGAAACAT
GCAATAATGCTAAGAGTAATTCCTCGACAGAAAAGGAGAATATGTTGGGGGAGGATTCAACTACTAAACAATCAAATCACTGTGAACTCCTGGCTTGCTC
TTCTCAGAGTGTACTGAAAGAGACGACTAAGAGTGTACCGAAAGAGACGACTAAGAGTATACTGAAAGAGACGACCAATAGTATACTGAAAGAAACGACT
AATACCAGTGGGGTCGATGCCAGGAAGCCACCAAAAAGGTTGCCAAACCAAGAAGAAGCAAAGGGAAACAAAGGAGGCAAAGTTCTCCCCAAGACAACTT
CATCAGACGAATCGCAGGTAACATTTCTATCTCGAATCGTTTTCCCCCCTGTTATGGATCGTGTCGATAGCCTGATGTTCGTTGGGTGTCGATTTCAGGT
TGTTAGAAGGTCTAAGCGATTGAGAAAATGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10025655 pacid=23145021 polypeptide=Lus10025655 locus=Lus10025655.g ID=Lus10025655.BGIv1.0 annot-version=v1.0
MDATPLKRVSEVGAELSRLTRPGKDTLIKLLRDGADALARIEHTSFLENSAKAEAHKKQATALKPLKESIVKRGLQRHTDRDVKLLVSVCVAELFRIQAP
ETPFADKYLGDVFKLLLGTFSELADVDSPYFGKRAGVLEAVARCECCVIMLDIGCGDLVVEMFNIFFSVARGHRGSSVERNILHIMSRILDEEPSQRLVD
IILQNLIKEEEGPSRLAVSIINKFHDTLKPSICRFLSSCFLERDAVESEVKESYHDILFQVYKHEPEMLLNVIPYLTEELRTDQVDVRLKAVTLLGKLLA
LRGDHFAQQHHSVLMEFKNRFSDKSVGVRLSALQCAKASYIANPRGKLSEEILGKSFSAFEGRLLDFSDDLSVRKKALVKLMELYRDYCSKCSERRMAIE
DHFEQIPPKVLMLWHIDSKEFRFQIVERVLGEDLFPANLSAEERSKHWVHIVSLFTPPHWKVLSSVLSQKNRLQTDMRTYLSLRMKAKGTGLEVMEGKIK
KLVINMSALFPDPSKAAECFNSLNQMKDCNIFSTLELLLKDSMITDVQVNRDKFQKVIGDKHPQHDFLQLLSSKCSFTIFNSEIIRCILVNISSNVLNSK
QKEASAKLLMAVISAFPFLLRGSEKEVQMLLGDNTLTAVLIEVLAKAGSHLSANLSDFYPFLRKVSLEGTRVQAKHAVSAFVSLAGSSEQAALSQLCEEL
VNSLRCGWNTSTVLQSLGCIMLHSVSASEIHGEEIRSYIFDIIFKLNQSDDPMCSGEMLNCTDSCKLKIFALKTLVRSVLPGQGSLLKWKINEMLGLLLK
MLRTGDTSDGVVSCASDKPYIRLAAAKSILQLSKRWDLHISHKIFQLTILMAKDPSPVVRRLFLYKTHKLLTNHAIPSRYACAFALAACGPCEDLREASF
KYMRDFIKYYVRISQFKQTSVGQRESPTDDPAFMVFFLIHILAHDDGFPHEDCQDEQVYARFCSPLFWAIQELTRSTNANGSLDFADYTILRLLSVFRAI
KRAEDAVDPQGTPKLQILADIGISIVNSLRHKGKSVSQTPGPVLLPSSLYKLSPVNLKRISSSADRQNFFSRLIPILKSEVVIPAETSNKRGRDCHEDKL
HSGDVEFSTSLSSELREPIHLQNLISTEVQATVAGDFSLGRLLEEALSPTSESDQLYSDDTSTEVNQGRACKFQKPTFSSSPILSGSAIATPSVLQCDVL
QQKSGGNVSSLQVTSGTADNSAEMELIRLDDETCNNAKSNSSTEKENMLGEDSTTKQSNHCELLACSSQSVLKETTKSVPKETTKSILKETTNSILKETT
NTSGVDARKPPKRLPNQEEAKGNKGGKVLPKTTSSDESQVTFLSRIVFPPVMDRVDSLMFVGCRFQVVRRSKRLRK
|
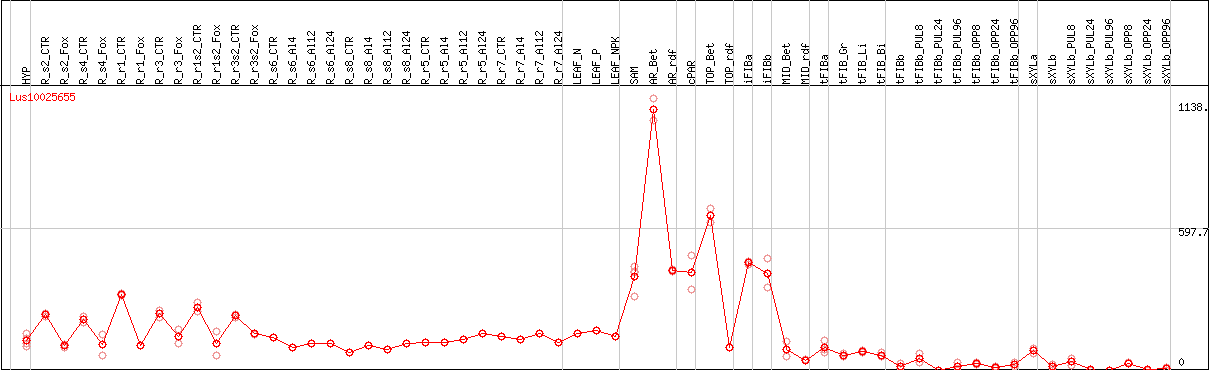
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025655 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
