External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G16240 292 / 4e-100
DELTA-TIP1, ATTIP2;1, AQP1, DELTA-TIP
delta tonoplast integral protein (.1)
AT4G17340 264 / 3e-89
TIP2;2, DELTA-TIP2
tonoplast intrinsic protein 2;2 (.1)
AT5G47450 262 / 3e-88
ATTIP2;3, DELTA-TIP3
DELTA-TONOPLAST INTRINSIC PROTEIN 3, ARABIDOPSIS THALIANA TONOPLAST INTRINSIC PROTEIN 2;3, tonoplast intrinsic protein 2;3 (.1)
AT2G36830 228 / 7e-75
TIP1;1, GAMMA-TIP1, GAMMA-TIP
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
AT3G26520 223 / 7e-73
TIP1;2, SITIP, GAMMA-TIP2, TIP2
SALT-STRESS INDUCIBLE TONOPLAST INTRINSIC PROTEIN, tonoplast intrinsic protein 2 (.1)
AT4G01470 209 / 2e-67
ATTIP1.3, GAMMA-TIP3, TIP1;3
tonoplast intrinsic protein 1;3 (.1)
AT2G25810 192 / 5e-61
TIP4;1
tonoplast intrinsic protein 4;1 (.1)
AT1G17810 188 / 4e-59
BETA-TIP
beta-tonoplast intrinsic protein (.1)
AT1G73190 184 / 1e-57
ALPHA-TIP, TIP3;1
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
AT3G47440 161 / 1e-48
TIP5;1
tonoplast intrinsic protein 5;1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038293
345 / 7e-121
AT3G16240 396 / 2e-141
delta tonoplast integral protein (.1)
Lus10004733
275 / 3e-93
AT5G47450 393 / 4e-140
DELTA-TONOPLAST INTRINSIC PROTEIN 3, ARABIDOPSIS THALIANA TONOPLAST INTRINSIC PROTEIN 2;3, tonoplast intrinsic protein 2;3 (.1)
Lus10007796
272 / 4e-92
AT5G47450 389 / 1e-138
DELTA-TONOPLAST INTRINSIC PROTEIN 3, ARABIDOPSIS THALIANA TONOPLAST INTRINSIC PROTEIN 2;3, tonoplast intrinsic protein 2;3 (.1)
Lus10022611
216 / 3e-70
AT2G36830 403 / 5e-144
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10014411
215 / 8e-70
AT2G36830 390 / 4e-139
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10005885
215 / 8e-70
AT4G01470 395 / 1e-140
tonoplast intrinsic protein 1;3 (.1)
Lus10021510
215 / 9e-70
AT2G36830 402 / 9e-144
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10023913
214 / 3e-69
AT2G36830 391 / 3e-139
TONOPLAST INTRINSIC PROTEIN 1;1, GAMMA TONOPLAST INTRINSIC PROTEIN 1, gamma tonoplast intrinsic protein (.1)
Lus10040863
210 / 7e-68
AT4G01470 394 / 2e-140
tonoplast intrinsic protein 1;3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00230
MIP
Major intrinsic protein
Representative CDS sequence
>Lus10025808 pacid=23177549 polypeptide=Lus10025808 locus=Lus10025808.g ID=Lus10025808.BGIv1.0 annot-version=v1.0
ATGGCGAAAATAGCATTCGGTAGATTTGATGACTCCTTCAGCTTCGGCTCAATCAAGGCCTACATCGCCGAGTTCATCTCTACCCTTCTCTTCGTTTTCG
CTGGGGTTGGATCCGCCATCGCTTTCAATAAGCTGACAGCAGATGCAGCACTCGACCCAGCTGGACTGGTCGCAGTTGCTGTCGGACACGCACTAGCCCT
CTTCGTCGCAGTCTCAGTCGGCGCCAACATCTCAGGTGGACATGTCAACCCAGCTGTCACCTTCGGATTGGCCGTCGGAGGCCAGATCACCATCCTCACT
GGCATCTTCTACTGGATTGCCCAGCTCGTCGGCTCCATCGTCGCTTGCTACCTCCTCAAAGTTGTCACCGGTGGCTTGGCTATTCCAACCCACGGCGTGG
CGGCGGGAGTGGGAGCATTGCAGGGAGTGGTGATGGAGATAATCATCACGTTCGGATTGGTGTACACGGTGTACGCGACCGCAGCTGATCCCAAGAAAGG
TTCGTTGGGAACCATTGCTCCGATTGCAATCGGGTTCGTGGTGGGTGCGAACATATTGGCAGCCGGCCCATTCTCAGGTGGATCAATGAACCCAGCAAGG
TCTTTCGGGCCGGCTGTGGCCAGTGCGGCTGTGGCCCGTGGGGGTTTCTCCGGAAACTGGATCTACTGGGTCGGGCCTCTGATCGGAGGTGGGCTTGCTG
GCATTATCTACCCCAACGTCTACATGTCCTCCGACCATGCTCCTCTGGCCAGTGACTACTGA
AA sequence
>Lus10025808 pacid=23177549 polypeptide=Lus10025808 locus=Lus10025808.g ID=Lus10025808.BGIv1.0 annot-version=v1.0
MAKIAFGRFDDSFSFGSIKAYIAEFISTLLFVFAGVGSAIAFNKLTADAALDPAGLVAVAVGHALALFVAVSVGANISGGHVNPAVTFGLAVGGQITILT
GIFYWIAQLVGSIVACYLLKVVTGGLAIPTHGVAAGVGALQGVVMEIIITFGLVYTVYATAADPKKGSLGTIAPIAIGFVVGANILAAGPFSGGSMNPAR
SFGPAVASAAVARGGFSGNWIYWVGPLIGGGLAGIIYPNVYMSSDHAPLASDY
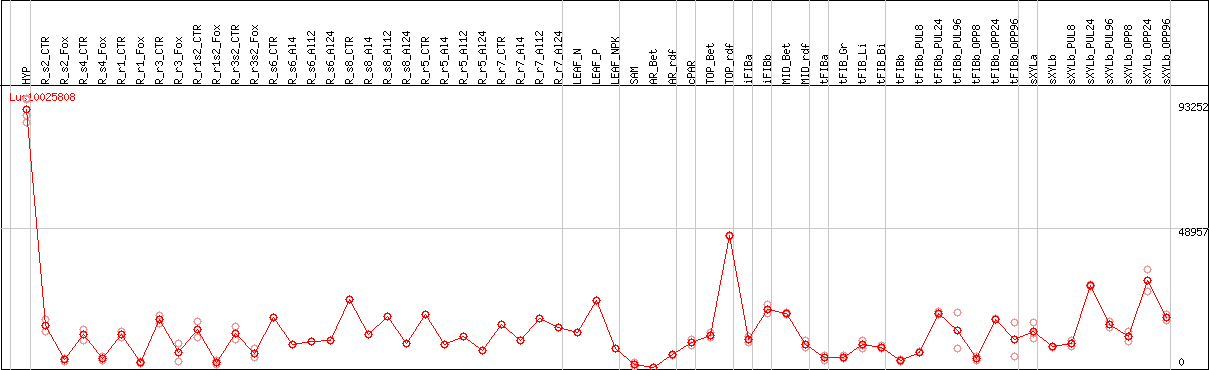
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025808 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.