External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G07370 375 / 4e-132
ATCHIP, CHIP
carboxyl terminus of HSC70-interacting protein (.1)
AT2G42810 71 / 8e-14
AtPP5, PP5.2, PP5, PAPP5
Arabidopsis thaliana protein phosphatase 5, protein phosphatase 5.2 (.1.2)
AT1G56440 63 / 5e-11
TPR5
tetratricopeptide repeat 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT2G45920 61 / 2e-10
U-box domain-containing protein (.1)
AT1G01670 61 / 3e-10
RING/U-box superfamily protein (.1)
AT3G17970 61 / 3e-10
ATTOC64-III
translocon at the outer membrane of chloroplasts 64-III (.1)
AT3G61390 59 / 9e-10
RING/U-box superfamily protein (.1.2)
AT1G01660 58 / 3e-09
RING/U-box superfamily protein (.1)
AT4G08320 56 / 8e-09
TPR8
tetratricopeptide repeat 8, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT4G11260 56 / 1e-08
RPR1, ETA3, EDM1, ATSGT1B, SGT1B
ENHANCER OF TIR1-1 AUXIN RESISTANCE 3, ENHANCED DOWNY MILDEW 1, phosphatase-related (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038219
564 / 0
AT3G07370 372 / 6e-131
carboxyl terminus of HSC70-interacting protein (.1)
Lus10020412
442 / 2e-158
AT3G07370 358 / 2e-125
carboxyl terminus of HSC70-interacting protein (.1)
Lus10009593
436 / 6e-156
AT3G07370 356 / 2e-124
carboxyl terminus of HSC70-interacting protein (.1)
Lus10034113
64 / 3e-11
AT3G59770 1913 / 0.0
ARABIDOPSIS THALIANA SUPPRESSOR OF ACTIN 9, sacI homology domain-containing protein / WW domain-containing protein (.1.2.3)
Lus10043471
60 / 5e-10
AT1G71020 682 / 0.0
ARM repeat superfamily protein (.1.2)
Lus10018004
60 / 6e-10
AT3G17970 711 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
Lus10031418
59 / 1e-09
AT1G56440 436 / 1e-150
tetratricopeptide repeat 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10018179
57 / 4e-09
AT4G08320 389 / 9e-134
tetratricopeptide repeat 8, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10025656
57 / 4e-09
AT4G08320 379 / 9e-130
tetratricopeptide repeat 8, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G248700
403 / 7e-143
AT3G07370 387 / 7e-137
carboxyl terminus of HSC70-interacting protein (.1)
Potri.005G177400
61 / 2e-10
AT4G08320 406 / 1e-139
tetratricopeptide repeat 8, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.005G014100
61 / 2e-10
AT1G56440 476 / 1e-165
tetratricopeptide repeat 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.014G141800
60 / 6e-10
AT2G42810 895 / 0.0
Arabidopsis thaliana protein phosphatase 5, protein phosphatase 5.2 (.1.2)
Potri.001G205300
58 / 2e-09
AT5G09420 724 / 0.0
outer membrane 64, ARABIDOPSIS THALIANA TRANSLOCON AT THE OUTER MEMBRANE OF CHLOROPLASTS 64-V, translocon at the outer membrane of chloroplasts 64-V (.1)
Potri.013G008400
57 / 6e-09
AT1G56440 471 / 1e-163
tetratricopeptide repeat 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.011G111500
57 / 7e-09
AT1G53300 707 / 0.0
tetratricopetide-repeat thioredoxin-like 1 (.1)
Potri.015G038600
56 / 1e-08
AT3G17970 747 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
Potri.001G216300
55 / 2e-08
AT3G11840 303 / 6e-99
plant U-box 24 (.1)
Potri.012G046900
55 / 2e-08
AT3G17970 744 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0229
RING
PF04564
U-box
U-box domain
CL0020
TPR
PF00515
TPR_1
Tetratricopeptide repeat
Representative CDS sequence
>Lus10025884 pacid=23177681 polypeptide=Lus10025884 locus=Lus10025884.g ID=Lus10025884.BGIv1.0 annot-version=v1.0
ATGGGGCAGGGAATTGTATTGACTCCGGCGAAACAAGCTGAGAGATTGAAAGAGAATGGAAATGCCTACTTCAAGAAAGAACGATTCGCTGCTGCCATCG
ACGCTTACACCGAGGCGATTGTGCTGTGCCCTAATGTTGCTGTGTACTGGACCAATCGAGCTCTTTGTCATCGTAAACGCAATAACTGGAAGAAAGTTGA
AGAGGATTGTCTTTCGGCGATCAAGCTTGATAACAATTCTTTTAAGGCTCACTATATGCTGGGACTTGCTTTACTCCAGAGGGAAGAGTACACAGAAGGA
CTCAAGGCATTGGAAAGAGCAATGGATCTTAGCAGGGATGCGAACCCACCAGGTTATATGGTGGAGGAGGTTTGGCAAGAGATTGCGAAAACAAAATACC
TGCTGTGGGAGCAAGCATCCACACAACGTTCATGGGAATTACAGAGCTTGAAAGAAGCTTGTGAAAGCGCTCTTAAAGAGAAGTACTTTCTCGATGATAA
TAAGAAAGATGGAGTTTCAGAGGAGGCAGCTGCCTTCCATGAGCAACAGTTGGATACGCTAAAGAAAGTGTTTGAGAAGGCAGCAGAAGATGACATCCCG
ACCGAGATACCAGATTATCTTTGTTGCAAAATCTCGCTCGATATCTTCCACGACCCTGTCATCACTCCAAGCGGGTTAACGTACGAAAAAGCAGTGATTC
TTGATCATCTTCAGAAGGTGGGGAGGTTTGATCCAATCACACGGGTAGAACTTGAACCATCTCAGTTGATCCCAAACCTGGCAATCAAGGAAGCCGTCCA
AACGTATCTTCAGAAACATGGTTGGGCATATAAAGCTGACTGA
AA sequence
>Lus10025884 pacid=23177681 polypeptide=Lus10025884 locus=Lus10025884.g ID=Lus10025884.BGIv1.0 annot-version=v1.0
MGQGIVLTPAKQAERLKENGNAYFKKERFAAAIDAYTEAIVLCPNVAVYWTNRALCHRKRNNWKKVEEDCLSAIKLDNNSFKAHYMLGLALLQREEYTEG
LKALERAMDLSRDANPPGYMVEEVWQEIAKTKYLLWEQASTQRSWELQSLKEACESALKEKYFLDDNKKDGVSEEAAAFHEQQLDTLKKVFEKAAEDDIP
TEIPDYLCCKISLDIFHDPVITPSGLTYEKAVILDHLQKVGRFDPITRVELEPSQLIPNLAIKEAVQTYLQKHGWAYKAD
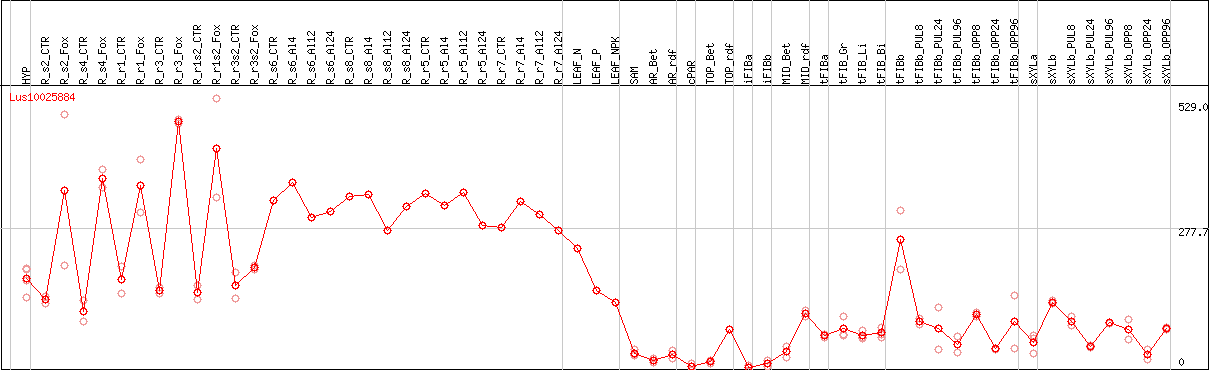
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025884 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.