External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G44230 167 / 5e-49
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G57430 139 / 3e-38
OTP84
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G23330 135 / 4e-37
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G13770 131 / 8e-36
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G14850 129 / 3e-35
MEF11, LOI1
lovastatin insensitive 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G08820 129 / 4e-35
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G26782 128 / 9e-35
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G24000 126 / 5e-34
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G02750 126 / 6e-34
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G04780 125 / 1e-33
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031989
142 / 2e-39
AT3G26782 865 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10020743
138 / 3e-38
AT1G04840 771 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10016387
134 / 2e-37
AT4G33990 717 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10033026
134 / 7e-37
AT1G08070 921 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010909
134 / 1e-36
AT5G66520 537 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010998
132 / 4e-36
AT4G16835 778 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10001220
131 / 1e-35
AT3G57430 1110 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019735
131 / 1e-35
AT4G33990 1015 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10004987
129 / 1e-35
AT3G46790 715 / 0.0
CHLORORESPIRATORY REDUCTION 2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G119000
214 / 3e-66
AT5G44230 792 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.016G051300
137 / 4e-38
AT3G13770 908 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.012G041200
136 / 1e-37
AT5G16860 1083 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G075800
136 / 2e-37
AT3G57430 1189 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G322100
134 / 9e-37
AT3G26782 897 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G215500
132 / 5e-36
AT3G63370 922 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 86, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G059400
132 / 6e-36
AT4G18750 1110 / 0.0
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G031600
131 / 7e-36
AT3G15930 784 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.018G040100
130 / 2e-35
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G231200
129 / 2e-35
AT5G66520 504 / 1e-172
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10025982 pacid=23173276 polypeptide=Lus10025982 locus=Lus10025982.g ID=Lus10025982.BGIv1.0 annot-version=v1.0
ATGTTAGTGTTGGAAATTGTCAAATTGGCTTATCACATTCATGCAAATCCAATTATAGCCGAAATTGCGGCAGCCCATCTGTTCGAGCTCGAACCGGACG
GTATTGGGAACTATATTCAGCTCTGCAGCACTTATGCAGCTGCTGAAAAATGGGACGATGTTTCTACGTTAAGGCGCATGATGAGAGGGAAAGGTCTCGA
CAAGAATCCTGGATATAGATGGATAGAAGCAGAAAAGGGCGTTATTCATAAGTTTGTTGCCCAGGACACGACCCATCCAAAGTCCAGGGAAATGAAGCAG
CTTCTTGAAGGTCTTCTAAACAAAATGGAGGCAAATGGGTACCAATGTAACCTGAGCACTATTCCGTACGGTGTGAGTGACGAGGAGAAGAGATCGGTTT
TGATGACTCGTAGCGAGAAATTGGCATTGGCCTTTGGGCTATTATGTTCCAAACCAGGCTCCACCATCAGGCTTATGAAGAACATCAGGATTTGCGACTG
CCATTTGTTCATGTGTGGAGCATCTTAA
AA sequence
>Lus10025982 pacid=23173276 polypeptide=Lus10025982 locus=Lus10025982.g ID=Lus10025982.BGIv1.0 annot-version=v1.0
MLVLEIVKLAYHIHANPIIAEIAAAHLFELEPDGIGNYIQLCSTYAAAEKWDDVSTLRRMMRGKGLDKNPGYRWIEAEKGVIHKFVAQDTTHPKSREMKQ
LLEGLLNKMEANGYQCNLSTIPYGVSDEEKRSVLMTRSEKLALAFGLLCSKPGSTIRLMKNIRICDCHLFMCGAS
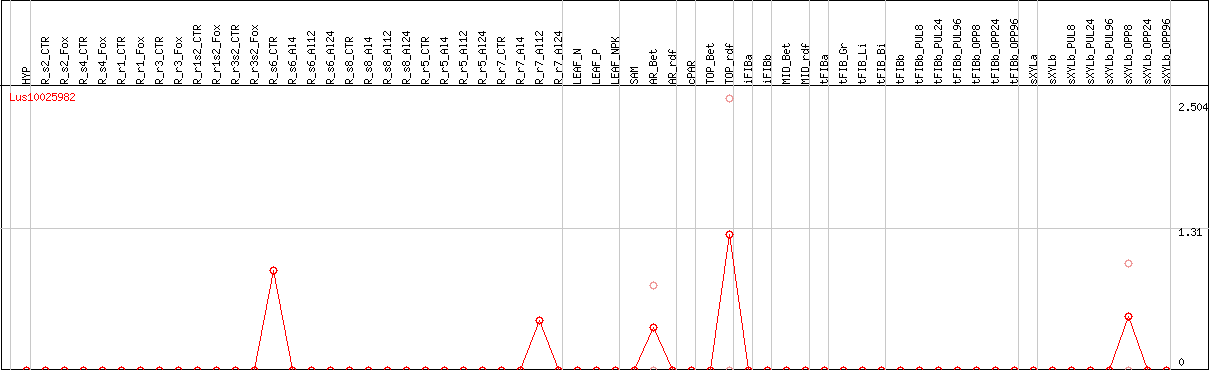
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10025982 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.