External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G09510 96 / 3e-24
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G66800 96 / 4e-24
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G35420 78 / 2e-17
TKPR1, DRL1
tetraketide alpha-pyrone reductase 1, dihydroflavonol 4-reductase-like1 (.1)
AT5G42800 76 / 1e-16
M318, TT3, DFR
dihydroflavonol 4-reductase (.1)
AT1G68540 62 / 8e-12
TKPR2, CCRL6
tetraketide alpha-pyrone reductase 2, cinnamoyl coA reductase-like 6, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT1G15950 62 / 9e-12
IRX4, ATCCR1, CCR1
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
AT1G25460 60 / 3e-11
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G58490 60 / 4e-11
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G33600 58 / 2e-10
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G27250 58 / 2e-10
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014363
88 / 3e-21
AT5G19440 420 / 3e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026070
85 / 6e-20
AT5G19440 419 / 4e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10002300
84 / 6e-20
AT5G19440 423 / 2e-149
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10019734
81 / 1e-18
AT1G09480 286 / 4e-96
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10002302
81 / 1e-18
AT5G19440 413 / 2e-145
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026072
82 / 2e-18
AT5G19440 339 / 1e-112
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10009955
80 / 4e-18
AT1G51410 548 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10006141
74 / 4e-16
AT4G35420 484 / 2e-173
tetraketide alpha-pyrone reductase 1, dihydroflavonol 4-reductase-like1 (.1)
Lus10041031
70 / 1e-14
AT5G42800 499 / 2e-178
dihydroflavonol 4-reductase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G057600
94 / 1e-23
AT5G19440 474 / 7e-170
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G256400
81 / 1e-18
AT5G19440 552 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.008G138600
79 / 8e-18
AT4G35420 535 / 0.0
tetraketide alpha-pyrone reductase 1, dihydroflavonol 4-reductase-like1 (.1)
Potri.005G229500
72 / 3e-15
AT5G42800 522 / 0.0
dihydroflavonol 4-reductase (.1)
Potri.008G120200
71 / 8e-15
AT1G68540 505 / 0.0
tetraketide alpha-pyrone reductase 2, cinnamoyl coA reductase-like 6, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G033600
70 / 1e-14
AT5G42800 523 / 0.0
dihydroflavonol 4-reductase (.1)
Potri.010G125400
68 / 7e-14
AT1G68540 428 / 1e-151
tetraketide alpha-pyrone reductase 2, cinnamoyl coA reductase-like 6, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.005G257700
66 / 3e-13
AT2G33590 400 / 4e-139
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G415500
62 / 1e-11
AT4G27250 422 / 3e-148
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.001G046100
62 / 1e-11
AT1G15950 511 / 0.0
IRREGULAR XYLEM 4, cinnamoyl coa reductase 1 (.1.2)
PFAM info
Representative CDS sequence
>Lus10026153 pacid=23173285 polypeptide=Lus10026153 locus=Lus10026153.g ID=Lus10026153.BGIv1.0 annot-version=v1.0
ATGAGCGGAGAAAGGGAAAGTGGTGTGTGTAACAGGAGCTTCAGGGTTCATAGCATCATGGTTGGTCAAGCTTCTGATTCACCATGGATACACTGTCAAC
GCTACTGTGACTCGAAGAAGACAGAGCATCTACTGAGGCTTGATGGAGCCAAGGAAAGACTGCATCTTTTCAAAGCAGACTTGCGGGAAGAAGGGTCTTT
TGATTCTGCTGTTCAAGGGTGTGATGGGGTTTTCCACACTGCTTCCCCTGTTTCCTTCTCTGCCGCTGATCCTCGGTCTGAAACAAAATCTACCTCATGG
AAGGGAGAATGTTCATATTGGTTGATCTTTCTTCTTGTTTGTAGATGTGATGAGATCGACAGTATCACAAAGGTGCCAGCTTTCAAGGCATCGAAGAAGA
AGGCAGAAGAAGAGTTGGGAATGAAGTTCATTGACACTATTGAGTGCCTGAAAGAAAAGGGCTTCTTACACTTCTGA
AA sequence
>Lus10026153 pacid=23173285 polypeptide=Lus10026153 locus=Lus10026153.g ID=Lus10026153.BGIv1.0 annot-version=v1.0
MSGERESGVCNRSFRVHSIMVGQASDSPWIHCQRYCDSKKTEHLLRLDGAKERLHLFKADLREEGSFDSAVQGCDGVFHTASPVSFSAADPRSETKSTSW
KGECSYWLIFLLVCRCDEIDSITKVPAFKASKKKAEEELGMKFIDTIECLKEKGFLHF
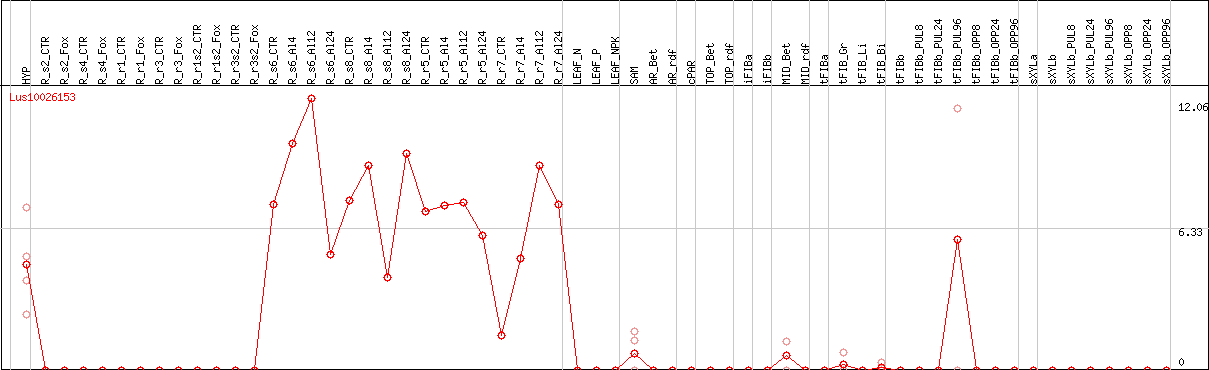
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026153 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.