External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G79400 238 / 2e-72
ATCHX2
cation/H+ exchanger 2, cation/H+ exchanger 2, cation/H+ exchanger 2 (.1)
AT1G16380 236 / 1e-71
ATCHX1
CATION EXCHANGER 1, Cation/hydrogen exchanger family protein (.1)
AT3G52080 162 / 7e-45
CHX28
cation/hydrogen exchanger 28 (.1)
AT2G13620 158 / 3e-43
ATCHX15
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
AT1G05580 142 / 1e-37
ATCHX23
cation/H+ exchanger 23, cation/H+ exchanger 23, cation/H+ exchanger 23 (.1), cation/H+ exchanger 23 (.2)
AT1G06970 133 / 1e-34
ATCHX14, CHX14
cation/hydrogen exchanger 14 (.1)
AT5G41610 133 / 2e-34
ATCHX18
cation/H+ exchanger 18, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 18, cation/H+ exchanger 18 (.1), cation/H+ exchanger 18 (.2)
AT2G31910 123 / 3e-31
ATCHX21
cation/H+ exchanger 21, cation/H+ exchanger 21, cation/H+ exchanger 21 (.1)
AT3G53720 120 / 3e-30
ATCHX20
cation/H+ exchanger 20, cation/H+ exchanger 20, cation/H+ exchanger 20 (.1)
AT2G30240 120 / 4e-30
ATCHX13
CATION/H+ EXCHANGER 13, Cation/hydrogen exchanger family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024131
402 / 3e-136
AT1G79400 580 / 0.0
cation/H+ exchanger 2, cation/H+ exchanger 2, cation/H+ exchanger 2 (.1)
Lus10024132
337 / 6e-110
AT1G16380 641 / 0.0
CATION EXCHANGER 1, Cation/hydrogen exchanger family protein (.1)
Lus10008666
314 / 1e-107
AT1G79400 162 / 3e-45
cation/H+ exchanger 2, cation/H+ exchanger 2, cation/H+ exchanger 2 (.1)
Lus10016403
159 / 1e-43
AT3G52080 856 / 0.0
cation/hydrogen exchanger 28 (.1)
Lus10019716
159 / 2e-43
AT3G52080 857 / 0.0
cation/hydrogen exchanger 28 (.1)
Lus10035312
157 / 1e-42
AT2G13620 1160 / 0.0
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Lus10030013
155 / 2e-42
AT2G13620 1160 / 0.0
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Lus10029506
154 / 8e-42
AT3G52080 850 / 0.0
cation/hydrogen exchanger 28 (.1)
Lus10032332
138 / 2e-36
AT4G23700 1030 / 0.0
cation/H+ exchanger 17, cation/H+ exchanger 17, cation/H+ exchanger 17 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G194200
337 / 2e-110
AT1G79400 625 / 0.0
cation/H+ exchanger 2, cation/H+ exchanger 2, cation/H+ exchanger 2 (.1)
Potri.003G194100
329 / 3e-107
AT1G79400 597 / 0.0
cation/H+ exchanger 2, cation/H+ exchanger 2, cation/H+ exchanger 2 (.1)
Potri.007G105200
168 / 6e-47
AT2G13620 1112 / 0.0
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Potri.009G059400
167 / 2e-46
AT3G52080 803 / 0.0
cation/hydrogen exchanger 28 (.1)
Potri.003G134900
145 / 1e-38
AT5G41610 989 / 0.0
cation/H+ exchanger 18, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 18, cation/H+ exchanger 18 (.1), cation/H+ exchanger 18 (.2)
Potri.001G264600
134 / 8e-35
AT3G52080 710 / 0.0
cation/hydrogen exchanger 28 (.1)
Potri.013G142301
132 / 2e-34
AT5G41610 1067 / 0.0
cation/H+ exchanger 18, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 18, cation/H+ exchanger 18 (.1), cation/H+ exchanger 18 (.2)
Potri.001G096320
132 / 2e-34
AT5G41610 1069 / 0.0
cation/H+ exchanger 18, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 18, cation/H+ exchanger 18 (.1), cation/H+ exchanger 18 (.2)
Potri.007G111400
131 / 8e-34
AT1G05580 645 / 0.0
cation/H+ exchanger 23, cation/H+ exchanger 23, cation/H+ exchanger 23 (.1), cation/H+ exchanger 23 (.2)
Potri.010G183000
125 / 7e-32
AT2G13620 474 / 8e-156
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
PFAM info
Representative CDS sequence
>Lus10026155 pacid=23173300 polypeptide=Lus10026155 locus=Lus10026155.g ID=Lus10026155.BGIv1.0 annot-version=v1.0
ATGTACCACCAGTTGAAAGACGGGGACCAATTCAGCGACGAAGAAGAGTACGGAGGGAACGACGTCGTCGAGATCAACGACGCGGTGGATTCATTCTCCG
CCGACACGAAATTCCTCATCCGCCAATCGAAGGTCGTCTCCACTTTCAACTCCTTGTTCGAGGACGTGTGCGGCGCCGTGGACGATCTCCGGATCTCGAT
CATTTTCTTAACGTTCCACAAGCACCAGCGGCTGGACGGGAAGCTGGAGAGAGGGAAGCCAGGGGTACGGACGACGAACCAGAAGGTTCTGCGGCAGGCG
GCGTGCTCGGTTGGGATATTCGTCGACAGAGGACAGACCGGATTCCAGATGCCGTCGCCGGACCATGTGCAGAACGTGGCGGTGATTTTCTTCGGCGGGC
CGGACGACAGGGACGCGGTAGCGTGTGGCAAGCGGATGGCGACGCATCCGTTCGTACGGATGACGCTGGTGAGATTTCTTCCGGCGGAGGAGGATGCTGA
GGCGGCGGAGAAAGGAGAATACGAAGGGAGGAGTAGCAGTCAGCGGAGCCAGGAGGAAGAAGTGTTGATGGAGATGCCGGAGGTGGAAGGGGAGATGGAT
AATGCCTGCATCAACGATTTCTACAACAGGTATGTAGCAGAAGGGAAAGTTGAATACGAAGAGATGGAAGTGAAAGATGGAAAGGAAACAGTAGAGGTAC
TGAAAGAGATTGGGGGAATGTATTCATTGTTCATAGTTGGGAAAGGAGGAGGTGGAGGTAGCTCTAGTTTGGCCGCCATGACCAGAGATATGAGCGATTG
GGAAGAATGTCCAGAACTTGGAACTGTGGGAGATCTGTTAGCTTCTACTGAGATCACCAGCATCAATGCTTCAGTATTGGTCATCCAACAGTATCGTCGT
TCTCCTTCTCAAATACGAGACTGA
AA sequence
>Lus10026155 pacid=23173300 polypeptide=Lus10026155 locus=Lus10026155.g ID=Lus10026155.BGIv1.0 annot-version=v1.0
MYHQLKDGDQFSDEEEYGGNDVVEINDAVDSFSADTKFLIRQSKVVSTFNSLFEDVCGAVDDLRISIIFLTFHKHQRLDGKLERGKPGVRTTNQKVLRQA
ACSVGIFVDRGQTGFQMPSPDHVQNVAVIFFGGPDDRDAVACGKRMATHPFVRMTLVRFLPAEEDAEAAEKGEYEGRSSSQRSQEEEVLMEMPEVEGEMD
NACINDFYNRYVAEGKVEYEEMEVKDGKETVEVLKEIGGMYSLFIVGKGGGGGSSSLAAMTRDMSDWEECPELGTVGDLLASTEITSINASVLVIQQYRR
SPSQIRD
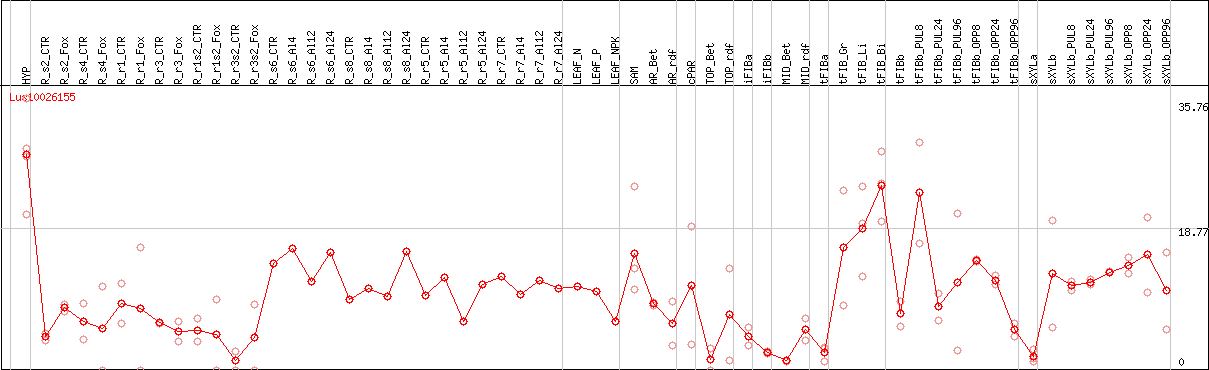
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026155 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.