External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G14570 86 / 2e-19
ATGSL4, ATGSL04
glucan synthase-like 4 (.1.2)
AT5G13000 78 / 1e-16
CALS3, ATGSL12
callose synthase 3, glucan synthase-like 12 (.1.2)
AT1G05570 73 / 1e-14
ATGSL6, ATGSL06, GSL6, CALS1
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
AT2G31960 70 / 9e-14
ATGSL3, ATGSL03
glucan synthase-like 3 (.1.2)
AT2G13680 68 / 5e-13
GLS2, ATGSL02, CALS5
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
AT3G59100 65 / 5e-12
ATGSL11
glucan synthase-like 11 (.1)
AT5G36870 62 / 5e-11
ATGSL9, ATGSL09
glucan synthase-like 9 (.1)
AT1G06490 60 / 2e-10
CalS7, ATGSL7, ATGSL07
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
AT4G03550 57 / 3e-09
ATGSL5, PMR4, GSL5, ATGSL05
POWDERY MILDEW RESISTANT 4, glucan synthase-like 5 (.1)
AT3G07160 50 / 5e-07
CALS9, ATGSL10
glucan synthase-like 10 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042478
138 / 3e-37
AT3G14570 2889 / 0.0
glucan synthase-like 4 (.1.2)
Lus10013744
77 / 4e-16
AT1G05570 2981 / 0.0
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
Lus10039199
77 / 4e-16
AT5G13000 3524 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Lus10037469
75 / 3e-15
AT1G05570 3143 / 0.0
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
Lus10003920
75 / 3e-15
AT1G05570 3231 / 0.0
GLUCAN SYNTHASE-LIKE 6, callose synthase 1 (.1.2)
Lus10020750
64 / 1e-11
AT1G06490 2003 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Lus10007327
64 / 1e-11
AT1G06490 1448 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Lus10020893
63 / 2e-11
AT2G13680 3037 / 0.0
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
Lus10001056
62 / 8e-11
AT1G06490 1596 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G095100
99 / 1e-23
AT3G14570 2863 / 0.0
glucan synthase-like 4 (.1.2)
Potri.001G230000
78 / 1e-16
AT5G13000 3157 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Potri.003G214200
78 / 1e-16
AT5G13000 3363 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Potri.001G012200
77 / 3e-16
AT5G13000 3492 / 0.0
callose synthase 3, glucan synthase-like 12 (.1.2)
Potri.005G058300
62 / 4e-11
AT2G13680 3190 / 0.0
ARABIDOPSIS THALIANA GLUCAN SYNTHASE-LIKE 2, callose synthase 5 (.1)
Potri.005G203500
61 / 7e-11
AT1G06490 2914 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Potri.002G058700
59 / 4e-10
AT1G06490 2961 / 0.0
callose synthase 7, Arabidopsis thaliana glucan synthase-like 7, glucan synthase-like 7 (.1)
Potri.013G131300
56 / 5e-09
AT4G03550 2752 / 0.0
POWDERY MILDEW RESISTANT 4, glucan synthase-like 5 (.1)
Potri.011G052400
49 / 1e-06
AT4G04970 2583 / 0.0
GLUCAN SYNTHASE LIKE-1, GLUCAN SYNTHASE LIKE 1, glucan synthase-like 1 (.1)
Potri.001G011900
42 / 0.0002
AT3G07160 2981 / 0.0
glucan synthase-like 10 (.1)
PFAM info
Representative CDS sequence
>Lus10026188 pacid=23156515 polypeptide=Lus10026188 locus=Lus10026188.g ID=Lus10026188.BGIv1.0 annot-version=v1.0
ATGGAAGCTGGTCTGGATAGTCCTTACATGGTTGCTGTTGCAATCTACATGACAACTAATGGAGTTGAGATGGTGTTATTTGTTGTTCCTGCTGTACGAA
AGTTCATTGAGATCTCAAATTGTCAGATTCAGAACTTTCTCTTGGTGGACACAGGTAAAGTTGGGCATAAAGGAGACCACAGACAAAAACAGCAACCAAG
ATTATATGCTGGTCGTGGGATGCAAGAAACCCAGGTTTCTGTTTTCAAGTACATTACTTTTCCTGCCATACCTTTCACTTTGATTTGTTTGCCTTCCCAG
TTTGAGTTTGCGAAATCTAAAAGTTGTTTTGTGCAGGGATTTGAAAGTGGAACAAGGACAAGAGCTGGCAGTCATGGTGGAACGACGAACAAACCCATCT
CCGGCGAGCAGGGGTTGGGGATAGACTTGGCTGTGGACATGGGAAGGCAACTGTTCAGTGCAAAATACCATTTGGGATTCAGGCTTTTCAAGACATTCCT
CTTTATTGCAGTTGTGGCCATCATAGTAACTCTATCACTAGCTGACAAGCTTTCGTTGAGGGATTTGATCGTCTGCTGTCTGGCATTTCTGCCAACAGGA
TGGGGACTTCTATTGATAGCACAAGCAGTGAGGCCGAAGATAGAGTAA
AA sequence
>Lus10026188 pacid=23156515 polypeptide=Lus10026188 locus=Lus10026188.g ID=Lus10026188.BGIv1.0 annot-version=v1.0
MEAGLDSPYMVAVAIYMTTNGVEMVLFVVPAVRKFIEISNCQIQNFLLVDTGKVGHKGDHRQKQQPRLYAGRGMQETQVSVFKYITFPAIPFTLICLPSQ
FEFAKSKSCFVQGFESGTRTRAGSHGGTTNKPISGEQGLGIDLAVDMGRQLFSAKYHLGFRLFKTFLFIAVVAIIVTLSLADKLSLRDLIVCCLAFLPTG
WGLLLIAQAVRPKIE
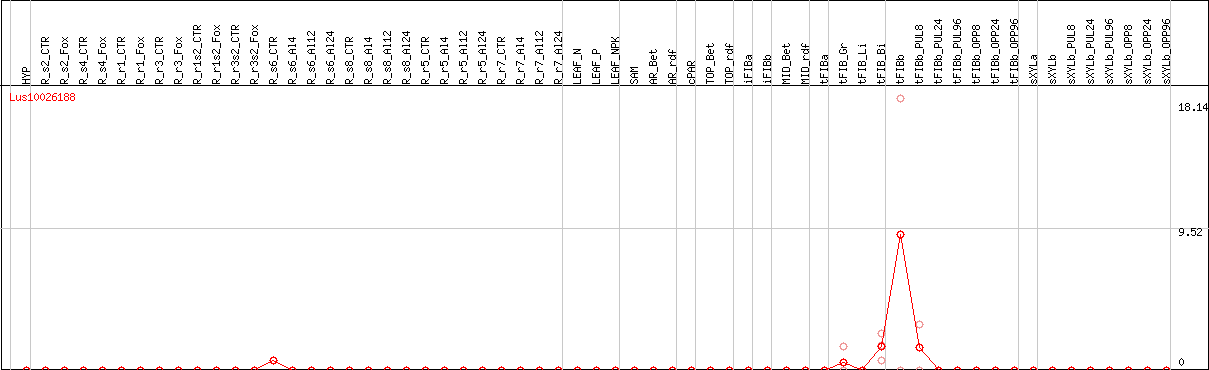
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026188 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.