Lus10026235 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10026235 pacid=23156620 polypeptide=Lus10026235 locus=Lus10026235.g ID=Lus10026235.BGIv1.0 annot-version=v1.0
ATGGCGGGGGGGGTTTGGGGGCGGCCCCCCTCATCTTTTTTAGATCTTTGCTTGTGGTTCTTCATCGCCATGACAGCTACTATTACGTCCGTCGTGGCAC
AGAGTGGCGGTAACTCGACGGTGGTTGCGGTGAACGTCGGGGTGATTCTGGACTTGAACAGGAGCAACTTCGTGGGACAGGTGGGGCTGAGCTGCGTGCA
AATGGCGATCTCCGATTTCTACGCTGCTCACCCTAACTACACCACCCGGATCACCATCAACACCAGGGATTCCGGCGGAGATGTCGTTACTGCGGCTGCT
TCAGCTCTGGATTTGATAAAGAACAAGCAAGTGGAAGCAATATTAGGACCAGAAACATCAATGCAGGCCAATTTCATCATAGGCATGGGGGAAAAGGCTC
GAGTCCCCGTCATCTCATTCTCTGCTTCAAGTCCAACCCTTGCTTCCACCCACAATCCCTATTTTTTACGTGCTACACCCAAGGACTCAACTCAAGTCGC
CGCCATTGCCGCCATTGTCAAGGCCTTTGGTTGGAGACAAGCTGTACCAATCTACACTGACAACGAATACGGAGAGGGTGTCATCCCTTACTTGATCGAC
GCCTTAGGGGAAGTCGATGCTCGTGTCCCATACCGCAGCAAGATCTCCCCATCCGCGACAGACGATCAAATAGGCCAGGAGCTCTACAAGCTTATGACCA
TGCAAACTAGAGTCTTCGTCGTACACATGCCGCCCCGTCTTGGGGCTAGGCTCTTCGCCAAGGCCAAAGAGATTGGAATGATGACATCAGGTTACGTCTG
GATCGTAACGGATGGAATCACCAATTTCTTTCCCTCCATGGACCGTTCTGGCCTCAACTTGATGCAGGGCGTTCTGGGTGTCAAACCTTACGTGCAGCTG
ACGAAACAAGTGGTCGATTTTCGGGTTAGGTGGAAGAGGAAATTCCAGCAAGAACATCCGGAGATTCTGGATTTTGACCTCAACAATTATGGGTTGTGGG
CGTATGATGCTACAGTGGCATTAGCCATGGCAGTTGAAAATACTTATAATGCTACTAATCCAAACTTGGGTTTTGCAAATGGCAATGTTTCAAGCAATGG
TTCTTCTTCTTCAACGACGGACCTTGGAAACTTCGGAGTGTCGAGAAACGGTCCAGATCTTGTCGAGAAGTTATCAAGAACAAGCTTCCAAGGACTATCA
GGTGAGTTTGTATTCAGTAATGGGGAGTTGAGAGATTCAGCTTACCAAATAGTGAATGTGAACGGTAATGGAGCAAGAGGGATTGGGTTCTGGAGCAGTG
AAAATGGGCTCACAAAAGAAATACTGAAATCAACCAGTACGGCACTACCAACTAAGGGAAGCCTTGGAACAGTGATATGGCCAGGGGATCCAGATTCTGT
TCCCAAAGGGTTTGAGATCCCAACAAATGGGAAGAAGCTGAGAATAGGAGTGCCTGTGAAAACTGGTTTCAACGAGTTCGTTTCCATCACAACCGATTCG
GCCACTAACGAGACAACTGTGGCCGGATACTCCATCAACATTTTCGAAGCTGTTGTCCAAGCATTGCCTTATGCTCTCCCCTTTGAGTATGTGCCTTTTG
CCACCGCGAATGGAGAGAGTGCTGGAACTTACAACGATCTGGTTTATCAGGTCTTCCTAGGGAAATTCGACGGCGTGGTAGCAGACACCACCATCATTGC
CAACAGATCACTGTACATCGACTTCACATTGCCGTACACAGAATCAGGAGTATCAATGATAGCACCAATCAAACCAATCCGAAACAAGAACGCTTGGGTA
TTCCTGAAACCACTGACGTGGGATCTTTGGGTGACGACGTTTCTCTTCTTCATTTTCATCGGGTTCGTGGTCTGGGTTTTGGAGCATAGAATCAACAAGG
ATTTCAGAGGACCTCCGTCCCATCAGGCCGGAACTAGCTTATGGTTCTCCTTCTCGGCCATGGTCTTCGCTCACAGAGAGACAGTGGTGAGCAACTTGGC
GAGGACGGTAGTGATCATATGGTGTTTTGTCGTGCTGATTTTGACGCAGAGTTACACGGCGAGTTTCACTAGCCTCTTAACAGTGCAGCAGCTCCAACCC
ACCGTAACGGACATAAATGAACTGCTGAGGAAAGGGGATTTCGTTGGGTATCAGGATGGGTCGTTTGTGTTGGGTCTCCTTAAGCGATTAGGATTCCCCG
AAGAGAAGCTTGTGCCTTATAAAAACTCTGATGACTGTGACTTGCTTTTCACCGAAGGAAGAATTGCTGCCGCATTCGACGAAGTGCCTTACATCAAGCT
CTTCTTAGCTAAGCACTGCTCCAAGTACACCACCGTCGACCCTACTTATAAGACTGACGGCTTTGGCTTCGTTTTCCCCAAAGGATCGCCGTTAGTTGCT
GACATTTCCAGGGCAGTTCTGAACGTAACAGAGGGGGACAAGATGAGCAAATTTGAGAAAGCATGGTTTGGTAAAAATAGCAGATGCCCAGAATCAAGTT
CCTCGGTTTCTCCAGGGAGCCAGCTGGGGCTAAACAGCTTCTGGAGCTTGTTCCTAATAGCTGGAGTTGCTGCAGTTTCAGCACTCTTCATCTTTACCGT
CACGTTTGTTTACCAACACAGGAATTATCTGCAGTTACCCCAGGAGGATAATTCAAATCTTTCATTATGGAGAAGGACTCTGAATCTTTTGAAGATATTT
GACAAAAAAGATCTCCAAAGCCATACTTTCAGGGAGGAGAGAAATGGCGGGGAGGTTCATCTTTCGGATTTGAATGTGTTAAGCCCTTCCGCATACTCGG
TTCAGACCGAGTTTCCTCAGGGGGATTCTTCTTCCTCGGAGTTGCATGACATTAACACAGCTGTTCAAGAAGTCGTTATAAATATTGACGATCCACATCC
GAATGGTGATGGTGAAGCACAGGTTGCTACTACAGATGAAGGAGTAACAAGGCCACAGGCAACCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10026235 pacid=23156620 polypeptide=Lus10026235 locus=Lus10026235.g ID=Lus10026235.BGIv1.0 annot-version=v1.0
MAGGVWGRPPSSFLDLCLWFFIAMTATITSVVAQSGGNSTVVAVNVGVILDLNRSNFVGQVGLSCVQMAISDFYAAHPNYTTRITINTRDSGGDVVTAAA
SALDLIKNKQVEAILGPETSMQANFIIGMGEKARVPVISFSASSPTLASTHNPYFLRATPKDSTQVAAIAAIVKAFGWRQAVPIYTDNEYGEGVIPYLID
ALGEVDARVPYRSKISPSATDDQIGQELYKLMTMQTRVFVVHMPPRLGARLFAKAKEIGMMTSGYVWIVTDGITNFFPSMDRSGLNLMQGVLGVKPYVQL
TKQVVDFRVRWKRKFQQEHPEILDFDLNNYGLWAYDATVALAMAVENTYNATNPNLGFANGNVSSNGSSSSTTDLGNFGVSRNGPDLVEKLSRTSFQGLS
GEFVFSNGELRDSAYQIVNVNGNGARGIGFWSSENGLTKEILKSTSTALPTKGSLGTVIWPGDPDSVPKGFEIPTNGKKLRIGVPVKTGFNEFVSITTDS
ATNETTVAGYSINIFEAVVQALPYALPFEYVPFATANGESAGTYNDLVYQVFLGKFDGVVADTTIIANRSLYIDFTLPYTESGVSMIAPIKPIRNKNAWV
FLKPLTWDLWVTTFLFFIFIGFVVWVLEHRINKDFRGPPSHQAGTSLWFSFSAMVFAHRETVVSNLARTVVIIWCFVVLILTQSYTASFTSLLTVQQLQP
TVTDINELLRKGDFVGYQDGSFVLGLLKRLGFPEEKLVPYKNSDDCDLLFTEGRIAAAFDEVPYIKLFLAKHCSKYTTVDPTYKTDGFGFVFPKGSPLVA
DISRAVLNVTEGDKMSKFEKAWFGKNSRCPESSSSVSPGSQLGLNSFWSLFLIAGVAAVSALFIFTVTFVYQHRNYLQLPQEDNSNLSLWRRTLNLLKIF
DKKDLQSHTFREERNGGEVHLSDLNVLSPSAYSVQTEFPQGDSSSSELHDINTAVQEVVINIDDPHPNGDGEAQVATTDEGVTRPQAT
|
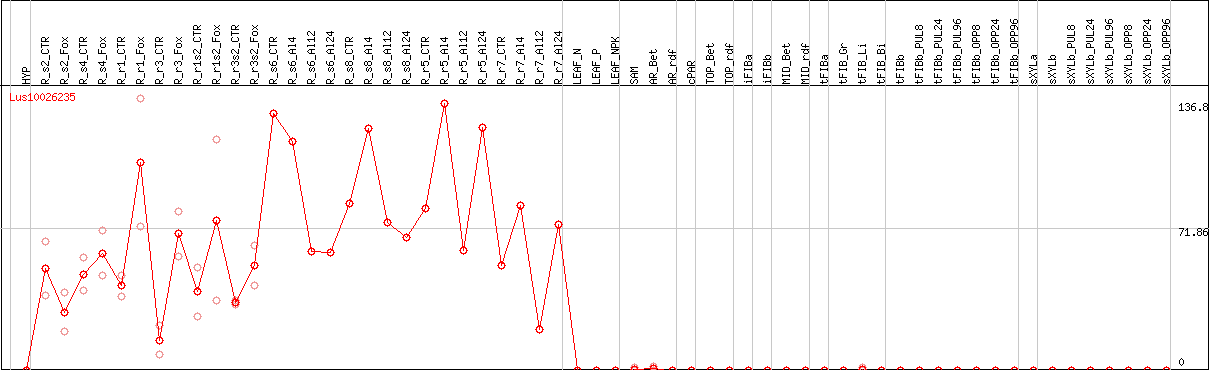
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026235 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
