External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G31820 75 / 4e-16
MAB4, ENP, NPY1
NAKED PINS IN YUC MUTANTS 1, MACCHI-BOU 4, ENHANCER OF PINOID, Phototropic-responsive NPH3 family protein (.1)
AT1G67900 67 / 2e-13
Phototropic-responsive NPH3 family protein (.1.2.3)
AT2G14820 62 / 1e-11
MEL3, NPY2
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
AT5G64330 61 / 6e-11
JK218, RPT3, NPH3
ROOT PHOTOTROPISM 3, NON-PHOTOTROPIC HYPOCOTYL 3, Phototropic-responsive NPH3 family protein (.1.2)
AT5G03250 60 / 9e-11
Phototropic-responsive NPH3 family protein (.1)
AT1G30440 59 / 1e-10
Phototropic-responsive NPH3 family protein (.1)
AT5G66560 59 / 2e-10
Phototropic-responsive NPH3 family protein (.1)
AT4G37590 58 / 4e-10
MEL1, NPY5
NAKED PINS IN YUC MUTANTS 5, MAB4/ENP/NPY1-LIKE 1, Phototropic-responsive NPH3 family protein (.1)
AT5G67440 58 / 4e-10
MEL2, NPY3
NAKED PINS IN YUC MUTANTS 3, MAB4/ENP/NPY1-LIKE 2, Phototropic-responsive NPH3 family protein (.1.2)
AT2G23050 57 / 1e-09
MEL4, NPY4
NAKED PINS IN YUC MUTANTS 4, MAB4/ENP/NPY1-LIKE 4, Phototropic-responsive NPH3 family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026255
221 / 4e-67
AT4G31820 557 / 0.0
NAKED PINS IN YUC MUTANTS 1, MACCHI-BOU 4, ENHANCER OF PINOID, Phototropic-responsive NPH3 family protein (.1)
Lus10042414
211 / 1e-65
AT4G31820 612 / 0.0
NAKED PINS IN YUC MUTANTS 1, MACCHI-BOU 4, ENHANCER OF PINOID, Phototropic-responsive NPH3 family protein (.1)
Lus10026702
65 / 2e-13
AT1G67900 207 / 5e-64
Phototropic-responsive NPH3 family protein (.1.2.3)
Lus10033796
65 / 1e-12
AT2G14820 735 / 0.0
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
Lus10000443
65 / 2e-12
AT1G67900 797 / 0.0
Phototropic-responsive NPH3 family protein (.1.2.3)
Lus10025192
64 / 3e-12
AT2G14820 248 / 2e-77
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
Lus10013799
63 / 6e-12
AT5G03250 672 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Lus10014632
63 / 7e-12
AT2G14820 741 / 0.0
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
Lus10026511
63 / 1e-11
AT5G03250 665 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G018600
120 / 4e-32
AT4G31820 630 / 0.0
NAKED PINS IN YUC MUTANTS 1, MACCHI-BOU 4, ENHANCER OF PINOID, Phototropic-responsive NPH3 family protein (.1)
Potri.006G264300
108 / 8e-28
AT4G31820 623 / 0.0
NAKED PINS IN YUC MUTANTS 1, MACCHI-BOU 4, ENHANCER OF PINOID, Phototropic-responsive NPH3 family protein (.1)
Potri.001G295600
65 / 2e-12
AT2G14820 771 / 0.0
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
Potri.008G186100
64 / 3e-12
AT1G67900 833 / 0.0
Phototropic-responsive NPH3 family protein (.1.2.3)
Potri.005G146400
64 / 3e-12
AT2G14820 671 / 0.0
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
Potri.009G089500
63 / 7e-12
AT2G14820 732 / 0.0
NAKED PINS IN YUC MUTANTS 2, MAB4/ENP/NPY1-LIKE 3, Phototropic-responsive NPH3 family protein (.1)
Potri.010G046800
62 / 1e-11
AT1G67900 861 / 0.0
Phototropic-responsive NPH3 family protein (.1.2.3)
Potri.008G038600
61 / 6e-11
AT5G03250 723 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Potri.016G090400
60 / 1e-10
AT5G03250 768 / 0.0
Phototropic-responsive NPH3 family protein (.1)
Potri.007G033900
59 / 2e-10
AT5G66560 789 / 0.0
Phototropic-responsive NPH3 family protein (.1)
PFAM info
Representative CDS sequence
>Lus10026256 pacid=23156415 polypeptide=Lus10026256 locus=Lus10026256.g ID=Lus10026256.BGIv1.0 annot-version=v1.0
ATGTGCAGCATGTTGGATGTTAAGAAGCTGACAATGAATGCTTCAATGCACGCAGCACAAAATGAGAGTCTCCCGCTTCGTTTGGTTGTACAAGTTCTCT
TCTTTGAGCAGGTCAGAGCAGCAGCAGCAGCAGCTAATGGTCAAAGCCTCAACCACAACCTGAATTACACTTCGAATGCGACGAAAAACACAGAGGAAGG
TTGCGAAGTCCTGGAGAAGCAAATGAGCGTGAAATTGAAGGTAGCAACTGTTGAGGCTGAGGAGGAGGAAGAGGAGGAGCAGAGGAATATGAGGCTGATG
AAGAAAAACAGCAGCAAGAATAGCAAAAGTGGATTGCAGTTGCTACCATCTCGTTCAAGGAGAATATTTGACAAGTTGTGGGGTGTTGGGAGGGGAGGGC
ATGTAGTAGAGAGCAAGAGCTCTGAGACATCTGGCAGCTCAGAGAGCCCACCTTCAATGGCTCCTGGAGATGCCAAGTCTTCTGGTTCATCTTCAAGGCA
CAGGAGACACTCCATCTCCTGA
AA sequence
>Lus10026256 pacid=23156415 polypeptide=Lus10026256 locus=Lus10026256.g ID=Lus10026256.BGIv1.0 annot-version=v1.0
MCSMLDVKKLTMNASMHAAQNESLPLRLVVQVLFFEQVRAAAAAANGQSLNHNLNYTSNATKNTEEGCEVLEKQMSVKLKVATVEAEEEEEEEQRNMRLM
KKNSSKNSKSGLQLLPSRSRRIFDKLWGVGRGGHVVESKSSETSGSSESPPSMAPGDAKSSGSSSRHRRHSIS
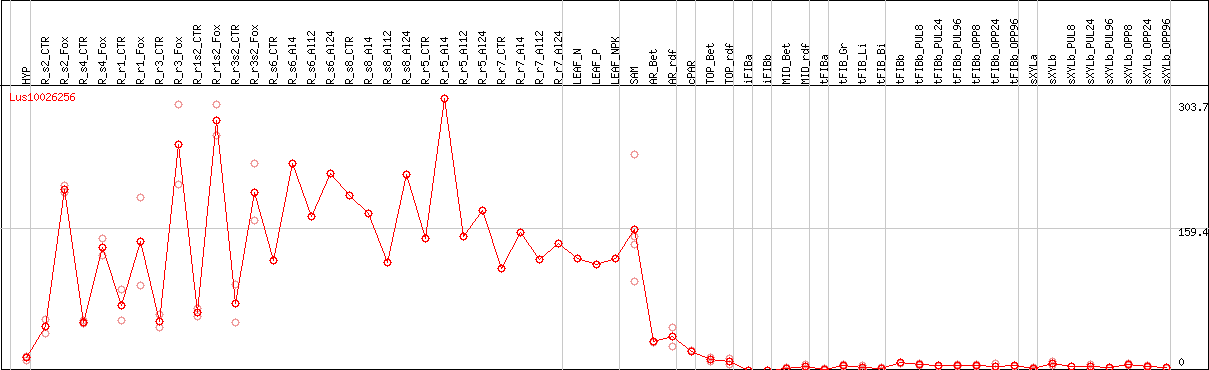
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026256 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.