External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G18830 135 / 4e-38
ATPMT5, AtPLT5
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
AT2G16120 125 / 3e-34
ATPMT1, AtPLT1
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 1, polyol/monosaccharide transporter 1 (.1)
AT2G16130 125 / 4e-34
ATPMT2, AtPLT2
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 2, polyol/monosaccharide transporter 2 (.1)
AT4G36670 118 / 1e-31
AtPMT6, AtPLT6
polyol/monosaccharide transporter 6, polyol transporter 6, Major facilitator superfamily protein (.1)
AT2G18480 118 / 1e-31
AtPLT3
Major facilitator superfamily protein (.1)
AT2G20780 93 / 1e-22
AtPLT4
Major facilitator superfamily protein (.1)
AT5G59250 69 / 2e-14
Major facilitator superfamily protein (.1)
AT4G16480 68 / 9e-14
ATINT4
inositol transporter 4 (.1)
AT1G30220 65 / 8e-13
ATINT2
ARABIDOPSIS THALIANA INOSITOL TRANSPORTER 2, inositol transporter 2 (.1)
AT2G35740 65 / 9e-13
ATINT3
nositol transporter 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026328
196 / 4e-62
AT3G18830 219 / 2e-66
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10042339
196 / 8e-61
AT3G18830 735 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10011288
179 / 2e-54
AT3G18830 721 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10040482
179 / 2e-54
AT3G18830 722 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10006971
140 / 6e-40
AT3G18830 631 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10011287
139 / 2e-39
AT3G18830 692 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10034623
136 / 3e-38
AT3G18830 652 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10035255
135 / 6e-38
AT3G18830 656 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Lus10041731
134 / 1e-37
AT4G36670 664 / 0.0
polyol/monosaccharide transporter 6, polyol transporter 6, Major facilitator superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G113600
165 / 5e-49
AT3G18830 717 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Potri.004G152300
159 / 1e-46
AT3G18830 625 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Potri.004G151725
150 / 8e-44
AT3G18830 640 / 0.0
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Potri.007G027900
124 / 7e-34
AT4G36670 638 / 0.0
polyol/monosaccharide transporter 6, polyol transporter 6, Major facilitator superfamily protein (.1)
Potri.006G158700
108 / 2e-28
AT3G18830 477 / 6e-165
ARABIDOPSIS THALIANA POLYOL/MONOSACCHARIDE TRANSPORTER 5, POLYOL TRANSPORTER 5, polyol/monosaccharide transporter 5 (.1)
Potri.013G135200
100 / 2e-25
AT2G20780 747 / 0.0
Major facilitator superfamily protein (.1)
Potri.004G039100
93 / 1e-22
AT2G20780 647 / 0.0
Major facilitator superfamily protein (.1)
Potri.016G010600
71 / 1e-14
AT4G16480 825 / 0.0
inositol transporter 4 (.1)
Potri.006G015200
67 / 3e-13
AT4G16480 811 / 0.0
inositol transporter 4 (.1)
Potri.001G244100
66 / 3e-13
AT5G59250 598 / 0.0
Major facilitator superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0015
MFS
PF00083
Sugar_tr
Sugar (and other) transporter
Representative CDS sequence
>Lus10026329 pacid=23156458 polypeptide=Lus10026329 locus=Lus10026329.g ID=Lus10026329.BGIv1.0 annot-version=v1.0
ATGGGGCCCATCACTTGGGTTTATAGCTCGGAGATTTTCCCCTTGAGGCTACGCGCTCAAGGGGCGAGTATGGGAGTGGCGGTGAACAGAGTGACGAGCG
GGGTTATATCGACGACGTTTATATCGATGTACAAAGGGATGACGATCAGCGGGGCATTCTTCTTGTTTGCCGGCATTGCTATGGTGGCGTTTGTGTTCTT
TTTCAGCTGTTATCCGGAGACTCAAGGGAGGACGTTGGAAGATATGGAGGTGCTGTTTGGTAGCTTCTTCGGATGGAGGTCTGCGGCTCTGGAGGTTGAG
AAGTTGAAGGAGAAAGAGGGCAAGAATGGTTGTCCAATGCTTCAGTATTGTCCTGGACTGATTAGTTTAATGATCATCCATGGCATGGACCAACCAAGCA
GGACAACAGTTCAGCAACAGCTGCCAGGCAACGCCAGGGAGTGCAGTTATAGTTTTGGATCCACTTCTTTCTAA
AA sequence
>Lus10026329 pacid=23156458 polypeptide=Lus10026329 locus=Lus10026329.g ID=Lus10026329.BGIv1.0 annot-version=v1.0
MGPITWVYSSEIFPLRLRAQGASMGVAVNRVTSGVISTTFISMYKGMTISGAFFLFAGIAMVAFVFFFSCYPETQGRTLEDMEVLFGSFFGWRSAALEVE
KLKEKEGKNGCPMLQYCPGLISLMIIHGMDQPSRTTVQQQLPGNARECSYSFGSTSF
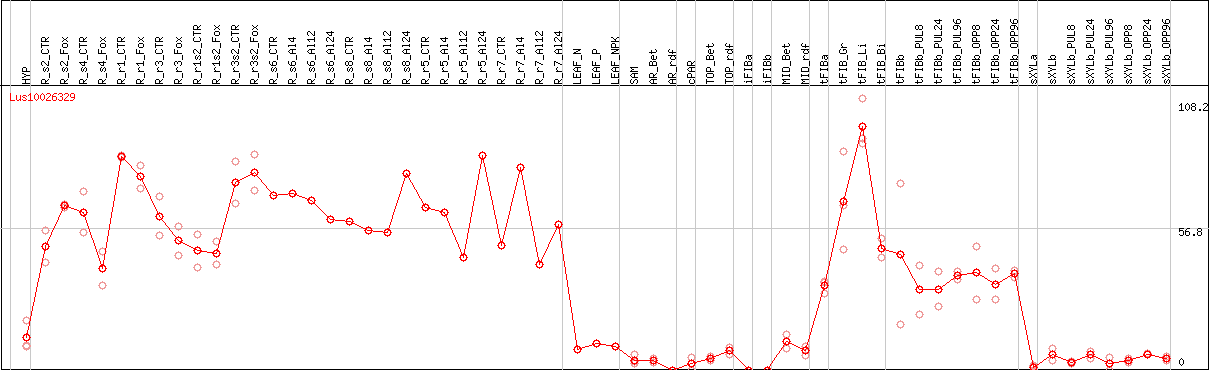
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026329 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.