Lus10026747 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10026747 pacid=23176573 polypeptide=Lus10026747 locus=Lus10026747.g ID=Lus10026747.BGIv1.0 annot-version=v1.0
ATGAGAAGGTGGGTTGCTAAGACTCTTCTTCAATCGCGTTCCCTTCCATTTCCAATCAGTCACTCGAAAACCAGGTTGCTCACAAGTTCTTCCTCCGACA
ATCTGGAAGGTTTGGTTGACCCGGATGACCTTTCTCTCTCAGGGCCAGAGTCAATTTCCAGTGAAGAGTTTACTCTTCTGCGGGATTCTTTATTGAACCC
TCATTCTTCTTCTTCTTCTTCTTCTTCTGCTGGTAAGTTCTCAGATGAAGCGTTGATGGTTGTTGATGTGATCTTGAGTGTTAATGATCAGTTTGGTAAA
GAAACCGTGAAGTTGCTCAGGCGGCATAGGGAGAAGTTAAGCGAGTCTTTGGTGATTGAGGTTCTGAATTTGCTCAATAGCCCTGAACTAGGTCTTAGAT
TCTTCATATGGGCTGGCCAGCAAATCGGATATTCCCACACTTTACCTGTGTACAATGCGCTACTAGAGTTGTTGCAGAGAAATAATAGTGCTGCTGGAGG
TAGAATACCAGAGCAGTTCCTGCTGCAAATCGAAGGTGGTGATGACAAGGAAGTTCTAGGGAAGTTACTTAACGTGTTGATTCGCAAATGTTGTCGCAAC
GGTTTGTGGAATGTTGCTCTGGAGGAGCTAGGTAGACTTAAGGATTATGGATACAAGCCGTCGAGATCGACTTACAATGCTTTGGTCCAAGTGTTCCTCA
GAGCTGAAAGGTTAGACACTGCTTCTCTGCTTCATAAAGAAATGTCAGGTTTAGGGTATGCAATGGATGGTTTTACTTTGGGATGTTTTGCTCGTTCTCT
TTGTAAAGCTGGGAAATGGAGAGAGGCATTGAAGTTAATTGAGGATGCGGAATTCAAACCTGATACTGTGCTTTTTACCAACATGATATCTGGATTGTGT
GAAGCCTCGCTCTTTGAAGTAGCTATGGATTTCTTAACAAGAATGCGAGTTTATTCTTGTATCCCTAATGTTCTGACTTATAGGATTTTGCTCTGTGGGT
GTTTGAGAAAGAAACAGCTTGGTAGATGTAAAAGAATTTTGAATATGATGATTACAGAAGGATGCTATCCAAGTCCAGCCATTTTTAACTCTTTAGTTAA
TGCTTATTGCAAATCGAGGGATTATGCTTACGCTTATAGGCTTATTAAGAAAATGATGGAATGTGGGTGCCAGCCGGGTTACATAGTCTACAACATATTT
ATTGGTGGTGTTTGTGGAAATCGGGAGTTGCCTGAGGCGGGTGCCCTTGAGTTGGCTGAAAAAGCATATGGGGAAATGCTCGATGCGGGAGTCACATTGA
ATAAGGTCAACGTTAGCAACCTTGCATGGTGTCTTTGTGGTGCTGGGAAATTCGAGGAGGCATATCGTCTCATTAGTGAAATGATGAGCAAGGGCTTTAT
ACCAGATACCAGTACATATTCCCAGGTCATTACTTGTCTCTGTAATTCGGCAAAAGTAGAAAAGGCCTTTCTATTATTTCAAGAAATGAGGAAGAATGGA
GTTCCGCCAGATGTGTACACATATACGCAATTGATCGACAGTTTTTGTAAGGTAGGTTTACTAGAACAGGCTCGCAGGTGGTTTGATGAAATGGATGAAA
ATGGATGTGCCCCTAATGTAGTGACTTACACAGCACTTATACATGCTTACCTTAAAGCAAGGAAGGTGTCCGATGCAAACCAAATTTTTGAGCTAATGTT
GTCCAAGGGCTCTGTTGCTAATGTCGTTACATATACAGCACTAATCGATGGACATTGTAAAGCAGGAGAGATCGAGACGGCTTGCAAAATATATGCTAGA
ATGAAACAGGACAATGTTGAAAATTCGGATGTGGAGTTGTATTTTGGGGTGATTGATACAGAGTCCGAGGAGCCAAATCATTTCACGTATGGAGCTTTGG
TTGACGGCCTATGCAAAGCTCATAAGATCAAAGAGGCCCGTGAGTTATTAGAAGCGATGTCAGTAAGTGGTTGTCAGCCTAATCATATCATCTATGATGC
ACTTTTAGATGGATGTTGCAAGATCGGTAAGCTAGATGAGGCGCAAGAGGTGTTTACCATGATGATAGAGCATGGATACAATCCAAATGTATATTCATAC
AGCTCTTTGATAGACAGGTTGTTTAAAGATAAGCGATTAGATCTCGCCTTGAAAGTCCTGTCCAAAATGCTTGAGAATTCTTGTGCTCCTAATGTTGTTA
TCTACACTGAAATGATTGACGGTCTATGCAAAGTTGGAAAAACTGATGAAGCATATAAACTTATGGTTATGATGGAAGAAAAGGAGTGTCATCCAAATGT
TGTCACCTATACTGCAATGATTGATGGGTTTGGGAAAGCTGGGAGACTTGACAAATGCCTTCAACTGTTTCAAAAGATGGCTTCCAAGGGCTGTGCTCCA
AATTTTGTCACTTATAAGGTTCTGATAAACCATTGTTGTGCAGCCGGCCTATTGGATGAGGCTCTTAAACTCTTGGAAGAGATGAAAGAGACACACTGGC
CAAAGCATGTAGCAGGTTATTGTAAAGTCATTGAAGGATTCAATCGAGAATTTATTGCTTCTCTGGATCTGTTGGATGAGTTTAGTGAAAAGGAAACAGT
ACCTGTTTCCCCTATTTACAGGGTTCTGATCGATAGTTTTGCCAAGGCCGGAAGACTGGAGAAAGCTTTAGAACTGTACAAAGAAGTCTCATCGCTCTTA
GCACCATCAGCAGCTAACCGGAATATGTATATTTCCTTAATAGAGGGCCTTGCACTTGCTAATAGGGTCGATAAGGCCTTTGATTTGCATGTCGACATGA
CTTTAAAAGGCATCATTCCTGAGGTAGGCACCGTCGTCCATGTGATCAAAGGTCTTTTGAGATTAAACAGGTGGGATCAAGTACTGCAGCTTTCGGATGG
AATATGCCAGGCGGATAATGAAAATGCATCCCCAGAGTTCTGCCATACCGAGAAGTCACTGAGAGTTGACAATGAGCAGTCCCAACCTTCCACTGAAAGG
ACTGAGGTGATGTCAATCCCTCCAACACGAATTTGGAAACAGTTGATCATGTGTCTTCTGGTGCAGAAAACCATCATGCTCTCCAAGAAACTAGCATTGA
TAAACAAGTTCGAATCAGGCATCTCTAAAAGCTGCAAAATCAAAGCAATCAGGCAGCTGGCGCAAATGATTTCACGAGGAGGCAACAAGCAGCAACCTTT
GGTTCTGGAAGCAAACATTGACACTCTGAGACAAGACGATGAGATAATCGAGCCTGCATCGTTGTATAATCTAGAACTCAAGAGTTTCAAGGGTGATTGT
ATGAGCCAGGTGCTATCTTTTCCCAAGGCTAGAATGTTCCTCACATTAGGTGATATTAATCAGATAAATGACTCAACAGCACAGCTTCTATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10026747 pacid=23176573 polypeptide=Lus10026747 locus=Lus10026747.g ID=Lus10026747.BGIv1.0 annot-version=v1.0
MRRWVAKTLLQSRSLPFPISHSKTRLLTSSSSDNLEGLVDPDDLSLSGPESISSEEFTLLRDSLLNPHSSSSSSSSAGKFSDEALMVVDVILSVNDQFGK
ETVKLLRRHREKLSESLVIEVLNLLNSPELGLRFFIWAGQQIGYSHTLPVYNALLELLQRNNSAAGGRIPEQFLLQIEGGDDKEVLGKLLNVLIRKCCRN
GLWNVALEELGRLKDYGYKPSRSTYNALVQVFLRAERLDTASLLHKEMSGLGYAMDGFTLGCFARSLCKAGKWREALKLIEDAEFKPDTVLFTNMISGLC
EASLFEVAMDFLTRMRVYSCIPNVLTYRILLCGCLRKKQLGRCKRILNMMITEGCYPSPAIFNSLVNAYCKSRDYAYAYRLIKKMMECGCQPGYIVYNIF
IGGVCGNRELPEAGALELAEKAYGEMLDAGVTLNKVNVSNLAWCLCGAGKFEEAYRLISEMMSKGFIPDTSTYSQVITCLCNSAKVEKAFLLFQEMRKNG
VPPDVYTYTQLIDSFCKVGLLEQARRWFDEMDENGCAPNVVTYTALIHAYLKARKVSDANQIFELMLSKGSVANVVTYTALIDGHCKAGEIETACKIYAR
MKQDNVENSDVELYFGVIDTESEEPNHFTYGALVDGLCKAHKIKEARELLEAMSVSGCQPNHIIYDALLDGCCKIGKLDEAQEVFTMMIEHGYNPNVYSY
SSLIDRLFKDKRLDLALKVLSKMLENSCAPNVVIYTEMIDGLCKVGKTDEAYKLMVMMEEKECHPNVVTYTAMIDGFGKAGRLDKCLQLFQKMASKGCAP
NFVTYKVLINHCCAAGLLDEALKLLEEMKETHWPKHVAGYCKVIEGFNREFIASLDLLDEFSEKETVPVSPIYRVLIDSFAKAGRLEKALELYKEVSSLL
APSAANRNMYISLIEGLALANRVDKAFDLHVDMTLKGIIPEVGTVVHVIKGLLRLNRWDQVLQLSDGICQADNENASPEFCHTEKSLRVDNEQSQPSTER
TEVMSIPPTRIWKQLIMCLLVQKTIMLSKKLALINKFESGISKSCKIKAIRQLAQMISRGGNKQQPLVLEANIDTLRQDDEIIEPASLYNLELKSFKGDC
MSQVLSFPKARMFLTLGDINQINDSTAQLL
|
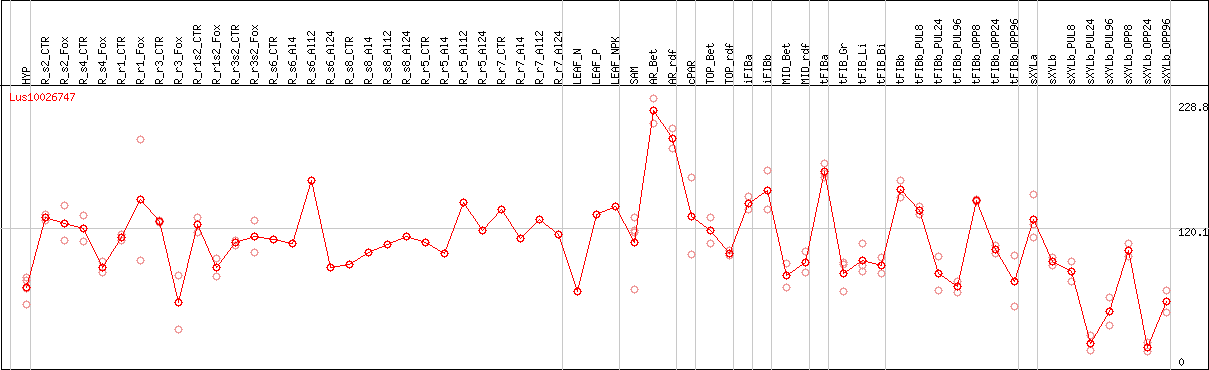
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026747 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
