External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G21090 248 / 9e-85
Leucine-rich repeat (LRR) family protein (.1)
AT3G43740 241 / 7e-82
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
AT1G71830 186 / 3e-56
ATSERK1, SERK1
somatic embryogenesis receptor-like kinase 1 (.1)
AT4G33430 182 / 6e-55
SERK3, RKS10, ELG, BAK1, ATSERK3, ATBAK1
RECEPTOR KINASES LIKE SERK 10, ELONGATED, SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE 3, BRI1-associated receptor kinase (.1.2)
AT1G34210 182 / 1e-54
ATSERK2, SERK2
somatic embryogenesis receptor-like kinase 2 (.1)
AT2G13790 172 / 4e-51
BAK7, BKK1, ATSERK4
BAK1-LIKE 1, BRI1- ASSOCIATED KINASE 7, somatic embryogenesis receptor-like kinase 4 (.1)
AT2G13800 154 / 3e-44
BAK8, ATSERK5
BRI1- ASSOCIATED KINASE 8, SOMATIC EMBRYOGENESIS RECEPTOR LIKE KINASE 5, somatic embryogenesis receptor-like kinase 5 (.1)
AT5G10290 115 / 5e-30
leucine-rich repeat transmembrane protein kinase family protein (.1)
AT3G25560 105 / 2e-26
NIK2
NSP-interacting kinase 2 (.1.2.3)
AT4G30520 102 / 2e-25
SARK
SENESCENCE-ASSOCIATED RECEPTOR-LIKE KINASE, Leucine-rich repeat protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002579
197 / 3e-64
AT5G21090 275 / 3e-94
Leucine-rich repeat (LRR) family protein (.1)
Lus10001807
189 / 3e-61
AT5G21090 227 / 9e-76
Leucine-rich repeat (LRR) family protein (.1)
Lus10002117
187 / 1e-60
AT3G43740 273 / 9e-94
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10013902
192 / 3e-57
AT1G49870 454 / 2e-145
unknown protein
Lus10020962
189 / 4e-57
AT4G33430 989 / 0.0
RECEPTOR KINASES LIKE SERK 10, ELONGATED, SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE 3, BRI1-associated receptor kinase (.1.2)
Lus10028716
186 / 2e-56
AT1G71830 1000 / 0.0
somatic embryogenesis receptor-like kinase 1 (.1)
Lus10004958
172 / 9e-55
AT1G34210 215 / 2e-66
somatic embryogenesis receptor-like kinase 2 (.1)
Lus10042755
167 / 1e-52
AT5G21090 199 / 8e-65
Leucine-rich repeat (LRR) family protein (.1)
Lus10000196
123 / 4e-37
AT5G21090 104 / 7e-30
Leucine-rich repeat (LRR) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G018800
253 / 2e-86
AT5G21090 354 / 6e-126
Leucine-rich repeat (LRR) family protein (.1)
Potri.001G219500
250 / 1e-85
AT5G21090 348 / 2e-123
Leucine-rich repeat (LRR) family protein (.1)
Potri.001G061600
202 / 1e-66
AT5G21090 272 / 2e-93
Leucine-rich repeat (LRR) family protein (.1)
Potri.003G166200
202 / 2e-66
AT5G21090 275 / 2e-94
Leucine-rich repeat (LRR) family protein (.1)
Potri.001G296500
195 / 7e-64
AT3G43740 278 / 1e-95
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.009G090700
190 / 7e-62
AT3G43740 270 / 2e-92
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.003G023000
186 / 5e-56
AT4G33430 961 / 0.0
RECEPTOR KINASES LIKE SERK 10, ELONGATED, SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE 3, BRI1-associated receptor kinase (.1.2)
Potri.001G206700
184 / 1e-55
AT4G33430 927 / 0.0
RECEPTOR KINASES LIKE SERK 10, ELONGATED, SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE 3, BRI1-associated receptor kinase (.1.2)
Potri.019G087700
182 / 2e-54
AT1G34210 1012 / 0.0
somatic embryogenesis receptor-like kinase 2 (.1)
Potri.007G082400
179 / 2e-53
AT1G34210 954 / 0.0
somatic embryogenesis receptor-like kinase 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0022
LRR
PF00560
LRR_1
Leucine Rich Repeat
CL0022
PF08263
LRRNT_2
Leucine rich repeat N-terminal domain
Representative CDS sequence
>Lus10026751 pacid=23176637 polypeptide=Lus10026751 locus=Lus10026751.g ID=Lus10026751.BGIv1.0 annot-version=v1.0
ATGGCGGCGGCTGCACCTCTCATCTTCGCTGCTGTTGCTCTAGCCCTGTGCTTCCCTCTCGCGTTTGCGAATTCGGAAGGAGATGCTCTCTACACACTTA
GGAGGAGCTTGTCCGATCCGGATAACGTCCTCCAGAGCTGGGATCCGACCTTAGTCAACCCTTGCACTTGGTTCCACATCACCTGCAACCAGGATAACCG
CGTTACCCGAGTGGACTTGGGGAATTCAAATTTGTCTGGGCATCTAGTGTCTGAGCTTGGGAAGCTAGAGCACTTACAGTATCTGGAGCTCTACAAAAAC
AACCTTCATGGAACTATCCCTGCTGAGCTTGGTAACTTGAAGAATCTGATCAGCTTGGACTTATACAACAACAACATAACCGGGACTATCCCTTCTTCGC
TTGGGGAATTGATTTCGCTTGGGAAATTGAAGAGTCTAGTCTTTTTAGATGTTACGAACAATGACTTGTGTGGAACTATTCCAACAACTGGACCATTCGA
GCATATCCCACTAAACAAATGA
AA sequence
>Lus10026751 pacid=23176637 polypeptide=Lus10026751 locus=Lus10026751.g ID=Lus10026751.BGIv1.0 annot-version=v1.0
MAAAAPLIFAAVALALCFPLAFANSEGDALYTLRRSLSDPDNVLQSWDPTLVNPCTWFHITCNQDNRVTRVDLGNSNLSGHLVSELGKLEHLQYLELYKN
NLHGTIPAELGNLKNLISLDLYNNNITGTIPSSLGELISLGKLKSLVFLDVTNNDLCGTIPTTGPFEHIPLNK
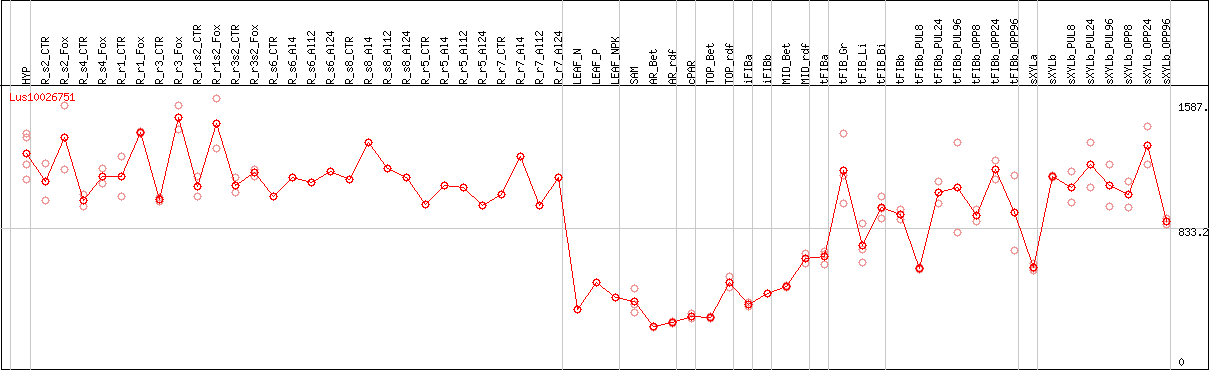
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026751 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.