External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G24740 149 / 1e-42
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
AT1G68140 147 / 3e-42
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
AT1G77770 141 / 1e-40
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
AT4G08460 135 / 5e-38
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
AT4G31410 134 / 2e-37
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
AT1G80220 127 / 3e-35
Protein of unknown function (DUF1644) (.1)
AT1G15430 122 / 2e-33
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
AT2G26050 91 / 1e-21
Protein of unknown function (DUF1644) (.1)
AT3G25910 92 / 4e-21
Protein of unknown function (DUF1644) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019436
165 / 6e-49
AT3G25910 277 / 3e-91
Protein of unknown function (DUF1644) (.1)
Lus10043292
162 / 9e-48
AT3G25910 282 / 2e-93
Protein of unknown function (DUF1644) (.1)
Lus10022809
154 / 2e-44
AT3G24740 351 / 1e-120
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
Lus10011876
152 / 4e-43
AT3G24740 358 / 5e-122
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
Lus10020151
149 / 5e-43
AT4G31410 363 / 1e-126
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
Lus10026950
147 / 2e-42
AT4G31410 363 / 2e-126
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
Lus10025356
139 / 7e-40
AT1G68140 282 / 3e-95
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Lus10020957
139 / 3e-39
AT1G68140 309 / 1e-104
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Lus10004940
140 / 5e-39
AT1G68140 319 / 4e-108
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G154300
171 / 5e-51
AT3G25910 152 / 8e-43
Protein of unknown function (DUF1644) (.1)
Potri.003G060800
170 / 1e-49
AT3G25910 188 / 3e-55
Protein of unknown function (DUF1644) (.1)
Potri.018G070500
163 / 4e-48
AT3G25910 246 / 2e-79
Protein of unknown function (DUF1644) (.1)
Potri.006G275400
147 / 2e-42
AT4G31410 311 / 4e-106
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
Potri.003G021800
148 / 3e-42
AT1G68140 315 / 2e-106
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Potri.018G006400
144 / 6e-41
AT4G31410 331 / 2e-113
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2)
Potri.001G208800
140 / 1e-39
AT1G68140 312 / 6e-106
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Potri.002G088700
135 / 1e-37
AT4G08460 245 / 2e-80
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Potri.005G172300
128 / 7e-35
AT4G08460 219 / 2e-70
Protein of unknown function (DUF1644) (.1), Protein of unknown function (DUF1644) (.2), Protein of unknown function (DUF1644) (.3)
Potri.001G173400
102 / 5e-25
AT3G25910 109 / 3e-27
Protein of unknown function (DUF1644) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0229
RING
PF07800
DUF1644
Protein of unknown function (DUF1644)
Representative CDS sequence
>Lus10026800 pacid=23176552 polypeptide=Lus10026800 locus=Lus10026800.g ID=Lus10026800.BGIv1.0 annot-version=v1.0
ATGAGTCCTATGATCAGACAAAAAAAGAAGGGATCATCAACATGGATGACGACGACGATAGAACACTGTTGCGAGAAGGAATGGGAAGAGATCAAGTGTC
CAATATGCATGGAACATCCTCACAACGCAATTCTGCTGCGTTGCTCATCGTACGAACTTGGATGTCGACCTTACCTGTGCAATACCAGCTCTCGACACTC
CAATTGTCTCGGAAAATTCTGCAATGCTTCCGAATCTTCCCTTTCTGAGCAAATTGGCAGCAAGGAAAACCTCATATGTCCACTTTGTCGTGGAGTGGTT
TACGACAGGACGGTGATCGACCCAGTCCGAAGTTACTTCAACACTAAATCGAGGACTTGCTCATCGCAGGATTGTGATTTTACTGGGAATTATTCTGAAC
TCGGAGCACATGCCAGATTTGAGCACCCTTTAAGCCGACCTATGCAAATCGATAGTGGGAGAGAGCGAGATTGGGATAGAATTGTACAAGCCCAAGAGTT
TGAGGACTTGGTTAGCACAATTGAACAACAAGATAATGATGAAGAAGTAGAGGTTCAGATTATAGTTCCAGAGAGCTACCTCCCTGCTTTGAGTAGTTTA
TTAGCAACTCTGTTTTATCAAACAAATCTTAATGATGACCAACATATGATTGTTGATGAATTGGAGTGGCCGGTTGCTACAACCCAACTTGGGGTTCAAA
GGAGTGATGCACGACGTCGTTTGGGTCGAAATTCGAGAATAAGAAACCTCAATGTTAGACATGATAACAATTTCTCTAGGAGAAGGGTTACATCAAATAC
ATTAAGCCAAAGGTACACTGCTAATAGCATCAACAACCATGATGCAAATAGGAGCTAG
AA sequence
>Lus10026800 pacid=23176552 polypeptide=Lus10026800 locus=Lus10026800.g ID=Lus10026800.BGIv1.0 annot-version=v1.0
MSPMIRQKKKGSSTWMTTTIEHCCEKEWEEIKCPICMEHPHNAILLRCSSYELGCRPYLCNTSSRHSNCLGKFCNASESSLSEQIGSKENLICPLCRGVV
YDRTVIDPVRSYFNTKSRTCSSQDCDFTGNYSELGAHARFEHPLSRPMQIDSGRERDWDRIVQAQEFEDLVSTIEQQDNDEEVEVQIIVPESYLPALSSL
LATLFYQTNLNDDQHMIVDELEWPVATTQLGVQRSDARRRLGRNSRIRNLNVRHDNNFSRRRVTSNTLSQRYTANSINNHDANRS
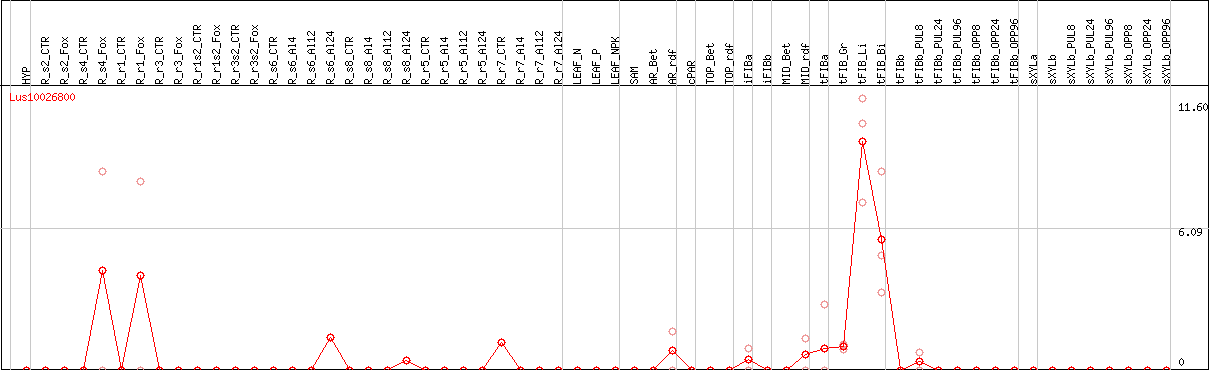
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.