External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G26820 156 / 2e-45
ATPP2-A3
phloem protein 2-A3 (.1)
AT1G33970 146 / 2e-43
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
AT1G33960 144 / 2e-41
AIG1
AVRRPT2-INDUCED GENE 1, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G09940 144 / 2e-41
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33950 141 / 8e-41
Avirulence induced gene (AIG1) family protein (.1), Avirulence induced gene (AIG1) family protein (.2)
AT1G33930 140 / 4e-40
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33830 132 / 2e-38
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G33910 133 / 5e-38
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G09930 131 / 7e-37
Avirulence induced gene (AIG1) family protein (.1)
AT4G09950 131 / 9e-37
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005300
219 / 5e-70
AT1G33970 321 / 7e-109
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10005299
184 / 3e-56
AT1G33970 339 / 5e-115
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10003732
149 / 1e-43
AT1G33970 210 / 1e-65
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10028023
138 / 3e-40
AT4G09940 181 / 2e-55
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10009482
115 / 1e-31
AT1G33970 148 / 9e-43
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009485
98 / 1e-25
AT1G33970 142 / 2e-42
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10009483
78 / 2e-18
AT1G33970 91 / 3e-23
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Lus10003507
74 / 2e-17
AT1G33930 82 / 3e-20
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10009484
69 / 2e-13
AT4G26220 255 / 1e-83
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G104800
196 / 6e-62
AT1G33970 314 / 3e-106
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Potri.019G077300
194 / 4e-61
AT1G33970 345 / 2e-118
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3.4.5)
Potri.009G131200
45 / 2e-05
AT3G16620 1129 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
Potri.004G171600
45 / 2e-05
AT3G16620 1128 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
Potri.009G131300
42 / 0.0002
AT3G16620 1111 / 0.0
ARABIDOPSIS THALIANA TRANSLOCON OUTER COMPLEX PROTEIN 120, translocon outer complex protein 120 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF04548
AIG1
AIG1 family
Representative CDS sequence
>Lus10026891 pacid=23142888 polypeptide=Lus10026891 locus=Lus10026891.g ID=Lus10026891.BGIv1.0 annot-version=v1.0
ATGGAAAATGAGAAGGAGTTGGTTACACCTTATAATACTGGTGTTGGAACCGTCGTTTTATGTGGACGTACTGGTAACAGGAAGAGTGCTACCGGGAACA
CCATTCTCGTCGAAAAGTGCTTCAAGTCTATGGTCAGTTCATCTGCTGTCACAACTTGCTACGATTTGCAGACGACTCTTATAGAAGATGGTCAAATCTT
AAACGTCATCGACACTCCTGCACTGCTTGGAAGTTGCGTTGAATCTGATTCTCTTGGCAAGGCAATCGTAAATATGGCCAGAAACAAGATCCCTGCAGTC
ATTCTTGTGTTATCATTAAGCAATCGCTTCACGGAAGAGGAAGAAGCAGCAATTCGTTGCCTCCAGTGTTTGTTCGGACGAAAGATAATGTACTACACGA
TTGTAGCGTTCACCGGTAGAGATAAACTTGAGGAGGAGGGTATTAAAACTTTGAATGAGTATTTATTTGGGTCGCAACTGTCCTCGGTCGTTAAAGAGTT
GTATGGATCGATGCCGGAAATTCTAGCACTTTGCGACAATCGGATGCTGCTGTTTGATAACCACACGAACGATGAAGTCAATAGAGTCGAACAGGTTAGG
CAGCTAATGACACTAGTAGACGGAATCGCTGTGCAGAATGGCGGGGAGTAA
AA sequence
>Lus10026891 pacid=23142888 polypeptide=Lus10026891 locus=Lus10026891.g ID=Lus10026891.BGIv1.0 annot-version=v1.0
MENEKELVTPYNTGVGTVVLCGRTGNRKSATGNTILVEKCFKSMVSSSAVTTCYDLQTTLIEDGQILNVIDTPALLGSCVESDSLGKAIVNMARNKIPAV
ILVLSLSNRFTEEEEAAIRCLQCLFGRKIMYYTIVAFTGRDKLEEEGIKTLNEYLFGSQLSSVVKELYGSMPEILALCDNRMLLFDNHTNDEVNRVEQVR
QLMTLVDGIAVQNGGE
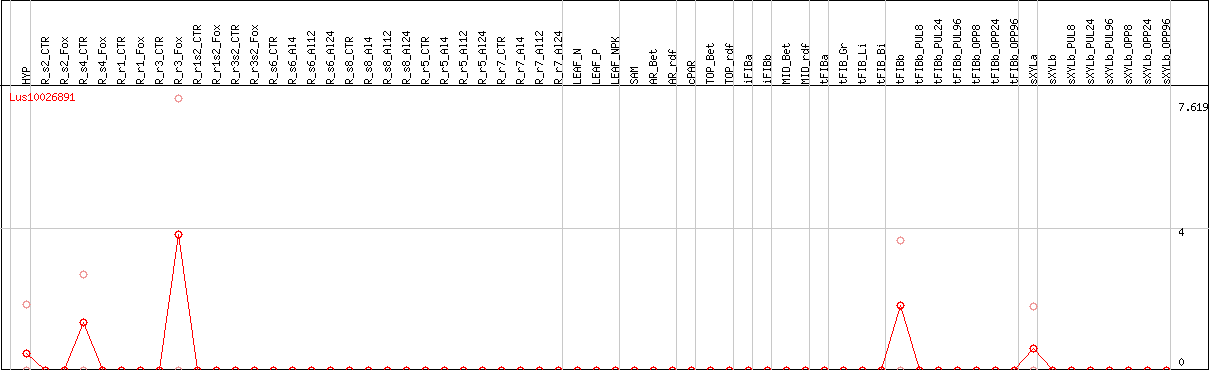
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10026891 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.