Lus10027175 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10027175 pacid=23145533 polypeptide=Lus10027175 locus=Lus10027175.g ID=Lus10027175.BGIv1.0 annot-version=v1.0
ATGCCAGCCGGAGTGATGGTTCCGGCGAGAAACATGGCATCATCGGCGGCGATTCTTAGGAGTAACGGCGGCACCGGCACCAGTAGTAGCTCTAGCGTTG
TTGGCCTTGGCGGGTTCGGCTCATCCACCACTCTTCCTATTCTTGGCCAGGGGAGCATGATGGACGACGTCGACGGGATGGTCGGTGGCGGCGGAGGGCA
CCACCACCACCACCAGCTTCAGTATTTACACCTAGACATGATGACCCACCACGCCACGCCGGAGAGCAACGACTTCCCCGGCGGAGGAGGAGAGTTTGAC
ATATCCGGCAACACAAAGTCCGGCGGGAGCGACAACCAAGACGGCGGCGGCGGATCTGGGGACGACCAGGAGCACCAGCAGCAACCGCCGAACAAGAAGA
AGCGGTACCACCGCCATACCCAGCACCAGATCCAGGAAATGGAAGCATTTTTCAAGGAATGTCCACACCCAGATGACAAGCAGAGAAAAGAGCTTAGCAG
GGAGCTAGGGCTCGAGCCTTTGCAAGTCAAGTTCTGGTTCCAGAACAAACGCACCCAGATGAAGACTCAACATGAGAGACTTGAGAACAATCAGCTGAGA
TCGGAGAACGACAAGCTTAGAGCCGATAACATGAGGTACCGTGAAGCCCTAACCAACACCTCCTGCCCGAACTGTGGTGGGCCCACCGCAGTCGGTGAGA
TGTCCTTCGACGAACACCATCTCAGGCTCGAGAATGCCCGACTTCGAGAGGAGATTGATCGGATATCAGGGATAGCTGCGAAGTACGTAGGGAAGCCGGT
GGTGAACTTCCCACTGCTGTCATCACCAATGGCACCACGTCCTGTCGAGCTACGGGTTGGGAATTTCGGAGGAGGTGAACATCAGCCTGCTGCCGCGGAC
GGAGGAGGAGGTGTGGATATGTATGGAGGTGGAGCAGGGGACTTGTTGAGGTCAATAAGTGGGCCCGCGGAGGCTGACAAGCCGATCATAATCGAGCTAG
CTGTTGCAGCCATGGAAGAGGTTGTTAGGATGGCTCAAATGGGTGAGCCCTTATGGATGACGAATGGTTTGGATGGTTCGGACCCGGTTTTGAACGAGGA
TGAGTACATTAGGACTTTCCCTCGTGGCATCGGACCCAAACCTAATGGGTTCAAGTCCGAGGCCTCGAGGGAGACCGCCGTTGTCATCATGAACCACGTC
AACCTCGTTGAGTACCTCATGGACGTGAACCAGTGGGCAGGTTTGTTCTCCGGGATAGTGTCTAGGGCAATGACATTGGAAGTGTTGTCAACGGGAGTTG
CGGGGAACTACAATGGTGCATTACAAGTGATGACAGCTGAACTCCAACTCCCGACTCCCCTTGTCCCCACGAGGGAGAGCTACTTTGCGAGGTACTGTAA
GCAACACTCCGACGGGACTTGGGCGGTGGTCGACGTTTCCTTGGACGACTTACGGCCGAGTCCGGTGGCAAGGTGCCGGAAGAGGCCTTCCGGTTGCTTG
ATTCAAGAAATGCCCAATGGTTACTCCAAGGTGACGTGGATTGAACACGTGGAAGTGGACGACAGAGGGGTCCACAATCTCTACAAGCAGATCGTCGGCT
CCGGCCACGCCTTCGGAGCAAAGCGGTGGGTGGCGACGCTGGACCGGCAGTGCGAGCGGCTGGCAAGCGCCATGGCTACAAACATCCCCACTAATGACGT
TGGAGTGATAACGAACCAAGAAGGGCGGAAGAGCATGATGAAGCTAGCAGAGAGGATGGTGGTCAGCTTCTGCGCCGGAGTCAGCGCTTCTACAGCTCAC
ACGTGGACGACTCTCTCCGGTACCGGAGCTGACGACGTCCGGGTCATGACCCGGAAGAGCATCGACGATCCGGGTCGTCCTCCCGGTATCGTGCTTAGTG
CCGCGACGTCATTCTGGATCCCTGTCGCGCCCAAACGTGTCTTTGACTTTCTCCGTGACGAAAACTCTCGTAGCGAGTGGGATATTTTGTCGAACGGAGG
GGCTGTACAAGAAAGGGCGCAAATAGCTAACGGCCGTGACACCGGCAACTGCGTATCTCTTCTTCGCGTTAACAGTGCGAATTCGAGCCAGGGGAACATG
CTGATACTGCAAGAGAGCTGCACGGATCAGACAGCGTCGTTCGTGATATATGCTCCGGTGGACATAGTGGCGATGAATGTGGTGCTCAACGGTGGGGATC
CGGATTATGTTGCTCTGCTTCCTTCAGGGTTCGCGATTCTTCCTGACGGCAACGCTGGTGGTGAAGCTGAAGGCGGCGGAGACGGAGGAGCAGGAGGAGG
AGGCGGTTCACTTCTAACTGTTGCATTTCAGATATTGGTGGATTCAGTTCCGACGGCGAAGTTATCTCTTGGATCAGTTGCTACTGTTAACAGTCTCATT
GCTTGCACCGTTGAGAGGATCAAGGCTGCTCTGTCCTGTGAGAATGCCGCATCTGCTAATCTTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10027175 pacid=23145533 polypeptide=Lus10027175 locus=Lus10027175.g ID=Lus10027175.BGIv1.0 annot-version=v1.0
MPAGVMVPARNMASSAAILRSNGGTGTSSSSSVVGLGGFGSSTTLPILGQGSMMDDVDGMVGGGGGHHHHHQLQYLHLDMMTHHATPESNDFPGGGGEFD
ISGNTKSGGSDNQDGGGGSGDDQEHQQQPPNKKKRYHRHTQHQIQEMEAFFKECPHPDDKQRKELSRELGLEPLQVKFWFQNKRTQMKTQHERLENNQLR
SENDKLRADNMRYREALTNTSCPNCGGPTAVGEMSFDEHHLRLENARLREEIDRISGIAAKYVGKPVVNFPLLSSPMAPRPVELRVGNFGGGEHQPAAAD
GGGGVDMYGGGAGDLLRSISGPAEADKPIIIELAVAAMEEVVRMAQMGEPLWMTNGLDGSDPVLNEDEYIRTFPRGIGPKPNGFKSEASRETAVVIMNHV
NLVEYLMDVNQWAGLFSGIVSRAMTLEVLSTGVAGNYNGALQVMTAELQLPTPLVPTRESYFARYCKQHSDGTWAVVDVSLDDLRPSPVARCRKRPSGCL
IQEMPNGYSKVTWIEHVEVDDRGVHNLYKQIVGSGHAFGAKRWVATLDRQCERLASAMATNIPTNDVGVITNQEGRKSMMKLAERMVVSFCAGVSASTAH
TWTTLSGTGADDVRVMTRKSIDDPGRPPGIVLSAATSFWIPVAPKRVFDFLRDENSRSEWDILSNGGAVQERAQIANGRDTGNCVSLLRVNSANSSQGNM
LILQESCTDQTASFVIYAPVDIVAMNVVLNGGDPDYVALLPSGFAILPDGNAGGEAEGGGDGGAGGGGGSLLTVAFQILVDSVPTAKLSLGSVATVNSLI
ACTVERIKAALSCENAASANL
|
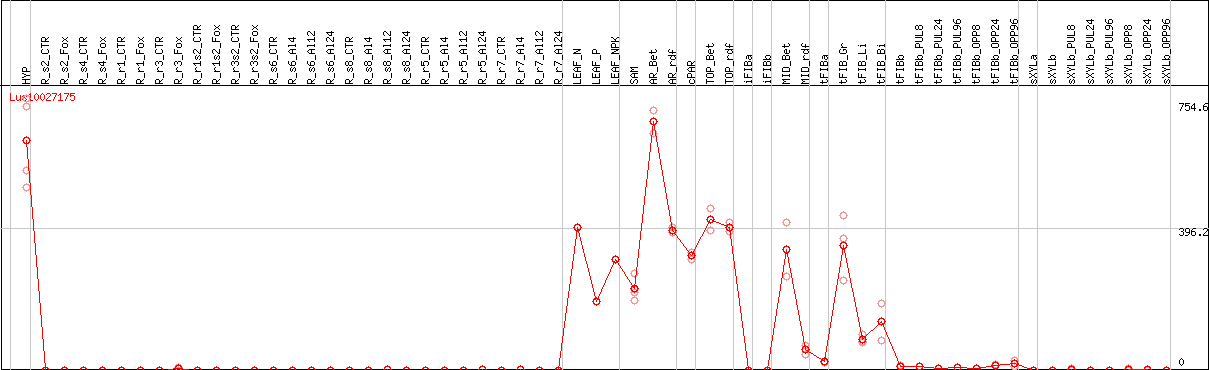
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027175 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
