Lus10027247 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10027247 pacid=23145508 polypeptide=Lus10027247 locus=Lus10027247.g ID=Lus10027247.BGIv1.0 annot-version=v1.0
ATGAAGAGCCTGTTGAAGAAGCTTCATATTACTAACCAGCAGGCGATTTCATCATCTTCAGAAGGGTCAAGTAATTCACCAAGAGCAAGTAAGTTTGGTG
GTGGTGATGTTTCGGTTTCGTCATCATCCCATGAGCACAACAAACCCTTTTCAGGTCTTTCAAATTGGCTGAGTACAGTTAAGATCAACAGGAAGAGTAC
TAGCCCTCCAAATTCGTCTCAAGGGGCGAAGATGAAGCCTAATCCAAATGATTCGGCCTCCAGTACTAGCTGTGGTGGCACCCCTAGAGGTGGTGCTGAG
ATTGATGAGGAGTATCAGGTGCAAATGGCCTTGGATCTCAGTACAATTGAGGATCCTGAGGCTGCTCAAATTGAGGCTGTCAAGCAGATAAACTCGGTGG
GCTCTGATGCTGATCTTGGGAAGAACACTCCTGTTGAGGTTCTTGCTTACCGTTACTGGAATTACAATGCTCTAAGCTACGACGACAGGATCGTGGACGG
TTTCTACGACTTGTACAGAATATCCACCGATTCCCAATCAGACAGGATGCCTTCCCTCATCGAGTTGCAAGGAGCTCTCGCCTCGGAAAGCATCACTTGG
GAGGCAGTGCTGGTTGATAGGACTACCGACGCGAATCTGCTGAAACTCGAACGTAAGTCGCAAGAAATTGCAAGTGAGACGAGGCCGGAGGAATCTCTGG
TTCTTGTGGACGGCAATCTGGTCAAGAAACTAGCTGATCTAGTTTCTGGCTACATGGGGGGACCTGTTGGTGACCCCGAGAACATGTCGGTAGCGTGGAG
ACGCCTTAGCTACAGTTTGAGGGCAACCCTTGGTAGCATGGTTTTGCCATTGGGGTCTCTAACTATTGGTCTGGCTCGCCATCGCACACTGATGTTCAAG
GTTTTAGCTGATAGTGTGGGCATACCGAGCCGGTTGGTGAAAGGGAATCAGTTCACGGGGTCAGACGATGTGGTCATCAACTTTGTGAAGCTTGATGATG
GAAGGGAGTACATTATCGATCTAATGGCGGATCCCGGGACTCTAATCCCTTCTGATGCAGCTGGATTACATCCGGAGCACGATGATCTTTCTTTCTCGGC
CAGTCCTTTGTCTAGAGACATTGAATCCTCTCATCTAGCCTCTTCTAGTAGTGGGGTGGTAAGTTCATTCGGAGAGACTTCGACCTCCGAAGTCGGAAAC
ACTATCGCTTCGAGTAATCAATGCAACGAAAGCGACAAAGAATCCCCGGATTTGTTAAAGAAGCAACAGAAGGAGATTGCTCCTAGTAGCAGCAGACAAA
ACTATACTTATACACACGTGAGATCGCCTTCGTGGACTGAGGGAATAAGCTCTCCAGCAGCACATAAGAAGAAAGTGAAAGACGTTTCGAGATACATGAT
AAACGCTGCAAAGGAGAATCCGCAGCTGGCTCAGAAACTTCACGATGTGCTGTTAGAGAGCGGTGTCGTTGCTCCGCCTAGCTTGTTTACCGAATTTGAG
CAGTTGAGGTTGAAGGATGTTGATTCCGGTCCTGCTCGATTTTTGCCTCCTCTGCCTCGTAAAGAAACTTCCATGCAGTCGAGGCTATCAACTCCTGAGA
CCATAGATGAGCCTAATCCGGCTCGATTCCTGCCACCTCTACCTCGGTGCAAACAGCATTCGGAAACAACTTCACCACAGGGTCAACTCGAGCCACTTAG
ATTCGACGAGTCTGAAGCAACTCCTGTGAGCTCTGCGAAAAAAGTTCCGGTTGCTGCTGCTGTGGCGGCAGCAGCAGCAGTCGTCGCTTCTTCGATGGTG
GTTGCTGCAGCAAGGCCAAACACCGAGTCGAGCCTCGAATTACCGATCGCTGCTGCTGCTGCAGCTGTCGTGGCATCAAGTGCAGCAGCTGTCGGTGTCA
AAATCGATGGAGATGGGGACCACCGGGGTCAAGAACATGATACGATTGCGGTGAACTCGGAGGGAGAGAGAACATCTGATAGATCGACGGGGAACGACAG
CTCCAGGTCGGATATCACTCTAGATGATGTTGCAGAGTGTGAGATCCCGTGGGAAGATATTACTTTGGGCGAACGAATTGGACTTGGTTCATATGGAGAA
GTTTACCATGGAGAATGGCATGGAACTGAAGTTGCTGTGAAGAGATTCCTAGATCAGGATATTTCAGGGGAATCTTTAATGGAGTTCAAAAGTGAGGTCC
AAATCATGAAGCGAGTTCGACATCCGAACGTGGTTCTTTTCATGGGAGCTGTGACGCGTGCTCCAAATCTTTCGATAGTTTCGGAGTTTCTTCCCAGAGG
TAGTTTGTACCGACTACTTCACCGTCCTAACAACCAGTTGGACGAGAGAAGACGACTCCGGATGGCACTTGATGCAGCTCGGGGGATGAATTACCTCCAT
AACTGCAGTCCAGTAATCGTTCATCGTGATTTGAAGTCTCCGAATCTCTTAGTTGATCGAAACTGGGTTGTGAAGGTATGCGATTTTGGTCTATCTAGAA
TGAAGCACAGCACATTCCTCTCCTCGAGATCAACTGCAGGAACTGCCGAGTGGATGGCTCCAGAAGTGCTGAGAAATGAACCTTCAGATGAGAAGTGCGA
TGTTTATAGCTACGGGGTTATACTGTGGGAGCTATGCACGATGCAACAACCGTGGGGAGGAATGAATCCGATGCAAGTCGTTGGTGCTGTGGGATTCCAG
CACCGACGACTCGAGATTCCAAACAATATGGATCCTGCTGTCGCAGATATTATTCAGAGTTGTTGGCAAACAGATCCTAAACTGAGGCCGACATTCGCAG
AGATCATGGCTGCATTGAAGCCACTGCAAAAGCCTATAACCAGCAGCTCGAGACGTGCAAGAACCGCATCTGTGACGAGTCCATTGCCAGAAGATGCTGC
AAAAGACCAGGCAGGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10027247 pacid=23145508 polypeptide=Lus10027247 locus=Lus10027247.g ID=Lus10027247.BGIv1.0 annot-version=v1.0
MKSLLKKLHITNQQAISSSSEGSSNSPRASKFGGGDVSVSSSSHEHNKPFSGLSNWLSTVKINRKSTSPPNSSQGAKMKPNPNDSASSTSCGGTPRGGAE
IDEEYQVQMALDLSTIEDPEAAQIEAVKQINSVGSDADLGKNTPVEVLAYRYWNYNALSYDDRIVDGFYDLYRISTDSQSDRMPSLIELQGALASESITW
EAVLVDRTTDANLLKLERKSQEIASETRPEESLVLVDGNLVKKLADLVSGYMGGPVGDPENMSVAWRRLSYSLRATLGSMVLPLGSLTIGLARHRTLMFK
VLADSVGIPSRLVKGNQFTGSDDVVINFVKLDDGREYIIDLMADPGTLIPSDAAGLHPEHDDLSFSASPLSRDIESSHLASSSSGVVSSFGETSTSEVGN
TIASSNQCNESDKESPDLLKKQQKEIAPSSSRQNYTYTHVRSPSWTEGISSPAAHKKKVKDVSRYMINAAKENPQLAQKLHDVLLESGVVAPPSLFTEFE
QLRLKDVDSGPARFLPPLPRKETSMQSRLSTPETIDEPNPARFLPPLPRCKQHSETTSPQGQLEPLRFDESEATPVSSAKKVPVAAAVAAAAAVVASSMV
VAAARPNTESSLELPIAAAAAAVVASSAAAVGVKIDGDGDHRGQEHDTIAVNSEGERTSDRSTGNDSSRSDITLDDVAECEIPWEDITLGERIGLGSYGE
VYHGEWHGTEVAVKRFLDQDISGESLMEFKSEVQIMKRVRHPNVVLFMGAVTRAPNLSIVSEFLPRGSLYRLLHRPNNQLDERRRLRMALDAARGMNYLH
NCSPVIVHRDLKSPNLLVDRNWVVKVCDFGLSRMKHSTFLSSRSTAGTAEWMAPEVLRNEPSDEKCDVYSYGVILWELCTMQQPWGGMNPMQVVGAVGFQ
HRRLEIPNNMDPAVADIIQSCWQTDPKLRPTFAEIMAALKPLQKPITSSSRRARTASVTSPLPEDAAKDQAG
|
DESeq2's median of ratios [FLAX]
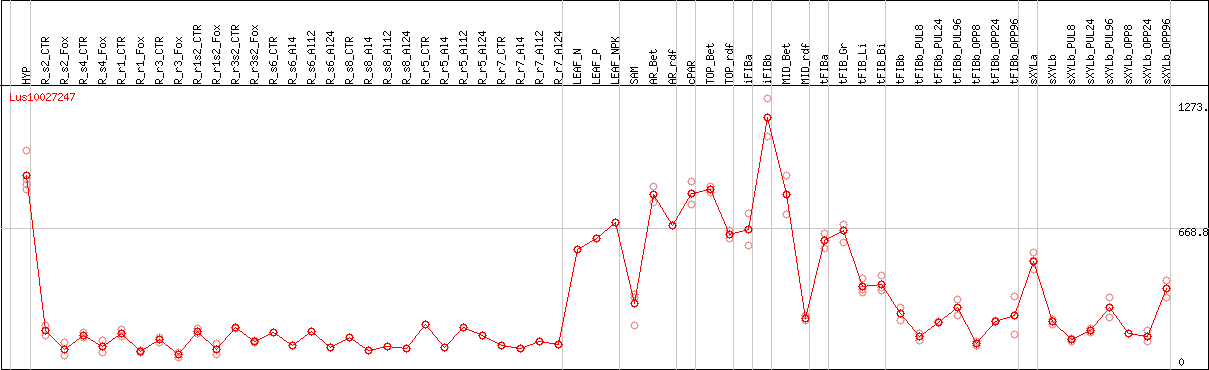
Coexpressed genes
Lus10027247 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
