External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G10200 260 / 2e-89
LIM
WLIM1, SF3
WLIM1, GATA type zinc finger transcription factor family protein (.1)
AT3G55770 193 / 6e-63
LIM
WLIM2b
WLIM2b, GATA type zinc finger transcription factor family protein (.1.2.3.4.5.6.7)
AT2G39900 189 / 3e-61
LIM
WLIM2a
WLIM2a, GATA type zinc finger transcription factor family protein (.1)
AT3G61230 169 / 3e-53
LIM
PLIM2c
PLIM2c, GATA type zinc finger transcription factor family protein (.1)
AT1G01780 167 / 2e-52
LIM
PLIM2b
PLIM2b, GATA type zinc finger transcription factor family protein (.1)
AT2G45800 166 / 7e-52
LIM
PLIM2a
PLIM2a, GATA type zinc finger transcription factor family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039010
360 / 5e-129
AT1G10200 285 / 2e-99
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10010455
305 / 3e-107
AT1G10200 277 / 3e-96
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10018674
266 / 2e-91
AT1G10200 323 / 4e-114
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10003822
264 / 1e-87
AT3G03220 409 / 4e-143
EXPANSIN 13, expansin A13 (.1)
Lus10007748
254 / 1e-86
AT1G10200 308 / 5e-108
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10004230
253 / 1e-86
AT1G10200 281 / 1e-97
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10042140
253 / 1e-86
AT1G10200 280 / 2e-97
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10043462
237 / 5e-80
AT1G10200 260 / 2e-89
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Lus10034123
218 / 3e-72
AT3G55770 244 / 1e-82
WLIM2b, GATA type zinc finger transcription factor family protein (.1.2.3.4.5.6.7)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G087200
295 / 5e-103
AT1G10200 316 / 2e-111
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Potri.001G292900
293 / 1e-102
AT1G10200 317 / 9e-112
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Potri.002G118000
265 / 4e-91
AT1G10200 325 / 7e-115
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Potri.014G015600
264 / 8e-91
AT1G10200 327 / 8e-116
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Potri.015G072400
244 / 6e-83
AT1G10200 277 / 5e-96
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Potri.012G077200
239 / 3e-81
AT1G10200 268 / 2e-92
WLIM1, GATA type zinc finger transcription factor family protein (.1)
Potri.008G063600
195 / 8e-64
AT3G55770 324 / 1e-114
WLIM2b, GATA type zinc finger transcription factor family protein (.1.2.3.4.5.6.7)
Potri.010G193800
195 / 1e-63
AT3G55770 323 / 2e-114
WLIM2b, GATA type zinc finger transcription factor family protein (.1.2.3.4.5.6.7)
Potri.003G124300
174 / 5e-55
AT3G61230 264 / 2e-90
PLIM2c, GATA type zinc finger transcription factor family protein (.1)
Potri.001G107100
172 / 2e-54
AT3G61230 256 / 6e-87
PLIM2c, GATA type zinc finger transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0167
Zn_Beta_Ribbon
PF00412
LIM
LIM domain
Representative CDS sequence
>Lus10027305 pacid=23145348 polypeptide=Lus10027305 locus=Lus10027305.g ID=Lus10027305.BGIv1.0 annot-version=v1.0
ATGGCCACATTTGCGATAACCAACCACAAATGCAAGGCTTGTGAGAAGACTGTCTATCTCGTCGACAAATTGGTCGCTGATACTCGTATCTACCACAAAT
CTTGTTTTAGATGCCACCACTGCAATAACACGCTCAAGCTGAGCAACTTCTGTTCTTTCGAAGGAGTCCTATACTGCCGGCCTCATTTTAACCAACTCTT
CAAAAGAACCGGTAGCCTTGACAAGAGCTTCGAAGGCACCCCCAAAGTTTCCAAGCCCGAAAAACCAATCGACACCGAGAATGCGAACAAGATGTCAAAT
CTCTTCGCAGGCACGAAGGAAAAGTGTAAGGAGTGCACAAAGACAGTGTATCCAACCGAGAGGGTGACAGTGAACGGAACATGGTACCATAGGAAGTGCT
TCAAGTGCAGTCATGGAGGATGCACAATTAGCCCATCAAACTACATTGCACATGAAGGCAAGTTGTATTGCAAGCACCACCATATCCAACTGTTCAAGGA
GAAAGGCAACTATAGCCAACTCGAAAACGATAATCATGACGACGACTGCGGCCGCCTCCCCAAAGCTGTTTGA
AA sequence
>Lus10027305 pacid=23145348 polypeptide=Lus10027305 locus=Lus10027305.g ID=Lus10027305.BGIv1.0 annot-version=v1.0
MATFAITNHKCKACEKTVYLVDKLVADTRIYHKSCFRCHHCNNTLKLSNFCSFEGVLYCRPHFNQLFKRTGSLDKSFEGTPKVSKPEKPIDTENANKMSN
LFAGTKEKCKECTKTVYPTERVTVNGTWYHRKCFKCSHGGCTISPSNYIAHEGKLYCKHHHIQLFKEKGNYSQLENDNHDDDCGRLPKAV
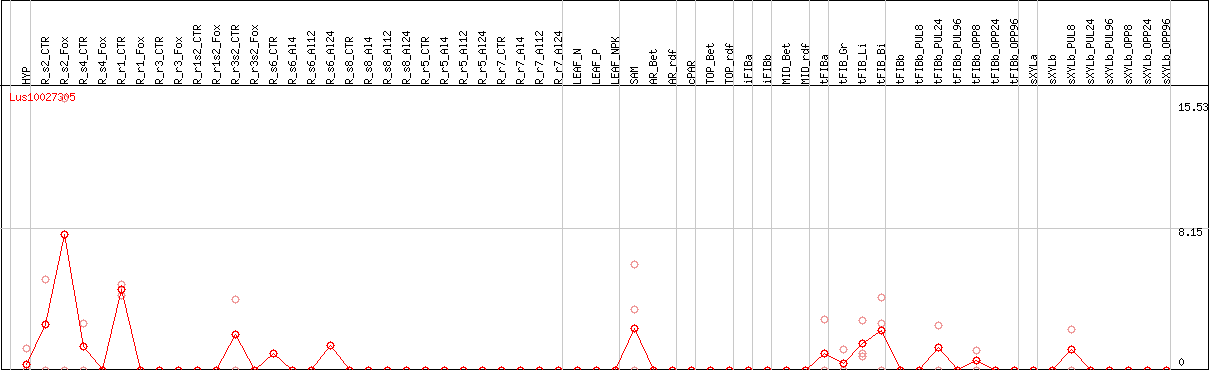
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027305 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.