Lus10027341 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10027341 pacid=23145566 polypeptide=Lus10027341 locus=Lus10027341.g ID=Lus10027341.BGIv1.0 annot-version=v1.0
ATGGCTTCTTCGACTGCCCAGATTCATGTTCTCGGAGGCATGGGGTTTGTTTCATCTGGGTCAAAGAAACCCAGCTTCTCTTTCGCCCCAAAGACCGTTT
TCCCGGGTCAAAGCTTGGGCGGTTCTAGAACGGCGGCGGTTTTGAGGTACAACGCGCGGCGGAGAAGCTACAGCAGCGGACGGTTGAGAGTGGCGTGTGA
GAAAGTGGTGGGAATCGATCTTGGTACCACCAATTCCGCTGTTGCCGCCATGGAAGGAGGGAAGCCGACTATTGTGACCAATGCGGAGGGGCAGAGGACT
ACACCGTCGGAGGTGGCGTATACGAAGAACGGTGATAGGCTGGTTGGTCAGATTGCGAAGCGGCAGGGAGTGGTGAACCCGGAGAACACTTTCTTCTCGG
TGAAGAGGTTCATCGGGAGGAAGATGTTGGAGGTGGATGAAGAGTCCAAACAGGTTTCTTATAGAGTGGTTAGGGACGACAATGGGAATGTTAAGCTGGA
TTGTCCAATTTTGGGTAAACAGTTTGCTGCTGAAGAGATTTCAGCTCAGGTTTTACGAAAACTCGTGGATGATGCTTCAAAATTTTTGAATGATAAAGTG
ACAAAAGCTGTGGTTACTGTACCTGCTTACTTTAACGACTCTCAAAGGACAGCAACGAAGGATGCTGGCCGAATTGCTGGGCTGGAAGTCCTCCGGATCA
TCAATGAGCCTACGGCTGCTTCACTGGCTTATGGATTTGAAAGGAAGAACAACGAGACCATCTTGGTGTTTGATCTTGGCGGTGGGACTTTCGATGTTTC
AGTTCTTGAGGTTGGTGATGGAGTCTTTGAAGTGCTCTCTACCTCTGGAGATACACATTTGGGTGGTGATGATTTTGACAAGAGAATTGTCGATTGGCTA
GCCGGGGACTTCAAGAGCAATGAAGGTATTGACCTTTTGAAAGACAAGCAGGCTCTCCAAAGGCTAACCGAGACAGCTGAGAAAGCCAAAATGGAGCTGT
CATCTTTGACTCAAGCAAACATCAGTTTGCCATTCATCACCGCAACCGCTGAGGGACCCAAGCACATCGAGACTACTCTTACGAGGGCTAAGTTTGAGGA
GTTATGCTCTGATCTCCTGGATAGGCTTAAAACTCCAGTTGAGAATTCCTTGAGGGATGCCAAACTTACCTTTAAGGACTTGGACGAAGTCATCCTTGTT
GGTGGCTCTACGCGTATACCTTCGGTGCAAGGACTTGTGAAGAAACTCACTGGGAAGGATCCTAATGTCACAGTAAATCCGGATGAAGTTGTTGCTTTGG
GCGCCGCAGTTCAGGCTGGTGTTTTGTCCGGTGATGTCAGCGACATTGTGCTGTTGGATGTCACGCCGCTATCAATTGGATTGGAGACCCTCGGAGGAGT
GATGACAAAGATCATTCCGAGAAACACCACCTTGCCATCATCCAAATCAGAGGTGTTCTCAACTGCTGCTGATGGGCAAACAAGTGTCGAGATCAACGTT
CTTCAAGGTGAGAGAGAGTTTGTTAGGGACAACAAGTCTCTAGGAAGCTTCCGCTTGGATGGCATCCCGCCAGCACCGCGTGGGGTCCCACAAATCGAAG
TTAAATTCGATATCGACGCTAATGGCATCCTCTCAGTCTCTGCTGCTGATAAGGGCACTGGGAAAAAACAAGACATTACAATCACTGGCGCCAGCACTCT
GCCTAGTGATGAGGTTTTACGAAAACTCGTGGATGATGCTTCAAAATTTTTGAATGATAAAGTGACAAAAGCTGTGGTTACTGTACCTGCTTACTTTAAC
GACTCTCAAAGGACAGCAACGAAGGATGCTGGCCGAATTGCTGGGCTGGAAGTCCTCCGGATCATCAATGAGCCTACGGCTGCTTCACTGGCTTATGGAT
TTGAAAGGAAGAACAACGAGACCATCTTGGTGTTTGATCTTGGCGGTGGGACTTTCGATGTTTCAGTTCTTGAGGTTGGTGATGGAGTCTTTGAAGTGCT
CTCTACCTCTGGAGATACACATTTGGGTGGTGATGATTTTGACAAGAGAATTGTCGATTGGCTAGCCGGGGACTTCAAGAGCAATGAAGGTATTGACCTT
TTGAAAGACAAGCAGGCTCTCCAAAGGCTAACCGAGACAGCTGAGAAAGCCAAAATGGAGCTGTCATCTTTGACTCAAGCAAACATCAGTTTGCCATTCA
TCACCGCAACCGCTGAGGGACCCAAGCACATCGAGACTACTCTTACGAGGGCTAAGTTTGAGGAGTTATGCTCTGATCTCCTGGATAGGCTTAAAACTCC
AGTTGAGAATTCCTTGAGGGATGCCAAACTTACCTTTAAGGACTTGGACGAAGTCATCCTTGTTGGTGGCTCTACGCGTATACCTTCGGTGCAAGGACTT
GTGAAGAAACTCACTGGGAAGGATCCTAATGTCACAGTAAATCCGGATGAAGTTGTTGCTTTGGGCGCCGCAGTTCAGGCTGGTGTTTTGTCCGGTGATG
TCAGCGACATTGTGCTGTTGGATGTCACGCCGCTATCAATTGGATTGGAGACCCTCGGAGGAGTGATGACAAAGATCATTCCGAGAAACACCACACTGCC
GTCATCCAAATCAGAGGTGTTCTCAACTGCTGCTGATGGGCAAACAAGTGTCGAGATCAACGTTCTTCAAGGTGAGAGAGAGTTTGTTAGGGACAACAAG
TCTCTAGGAAGCTTCCGCTTGGATGGCATCCCGCCAGCACCGCGTGGGGTCCCACAGATCGAAGTTAAATTCGATATCGACGCCAATGGCATCCTCTCAG
TCAGTGCTGCTGATAAGGGCACTGGGAAAAAGCAAGACATTACAATCACTGGTGCCAGCACTCTGCCAAGTGATGAGGTTGAGAGGATGGTCCAAGAAGC
GGAGAGGTTTTCGAAGGAGGACAAGGAGAAGAGGGACGCCATTGACACGAAGAACCAAGCTGATTCTGTTGTTTATCAAACCGAGAAGCAGCTCAAGGAG
CTCGGAGACAAAGTTCCGGGCGAGGTGAAAGAAAAGGTTGAGTCTAAATTGCAAGAGCTGAGGGATGCCATTGCTGGAGGATCAACTCAAACAATGAAAG
ATGCGATGACAACTCTCAATCAGGAAGTCATGCAACTCGGCCAGTCCCTTTACAACCAACAAGGTTCGGGTGCGGGTGAAGGTCCAACAACTCCGGGGAG
TGAATCTGGGCCATCGTCGTCTTCCGATTCTTCTAGCAAGGGACCCGATGGAGATGTCATTGATGCCGATTTCACAGACAGCAAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10027341 pacid=23145566 polypeptide=Lus10027341 locus=Lus10027341.g ID=Lus10027341.BGIv1.0 annot-version=v1.0
MASSTAQIHVLGGMGFVSSGSKKPSFSFAPKTVFPGQSLGGSRTAAVLRYNARRRSYSSGRLRVACEKVVGIDLGTTNSAVAAMEGGKPTIVTNAEGQRT
TPSEVAYTKNGDRLVGQIAKRQGVVNPENTFFSVKRFIGRKMLEVDEESKQVSYRVVRDDNGNVKLDCPILGKQFAAEEISAQVLRKLVDDASKFLNDKV
TKAVVTVPAYFNDSQRTATKDAGRIAGLEVLRIINEPTAASLAYGFERKNNETILVFDLGGGTFDVSVLEVGDGVFEVLSTSGDTHLGGDDFDKRIVDWL
AGDFKSNEGIDLLKDKQALQRLTETAEKAKMELSSLTQANISLPFITATAEGPKHIETTLTRAKFEELCSDLLDRLKTPVENSLRDAKLTFKDLDEVILV
GGSTRIPSVQGLVKKLTGKDPNVTVNPDEVVALGAAVQAGVLSGDVSDIVLLDVTPLSIGLETLGGVMTKIIPRNTTLPSSKSEVFSTAADGQTSVEINV
LQGEREFVRDNKSLGSFRLDGIPPAPRGVPQIEVKFDIDANGILSVSAADKGTGKKQDITITGASTLPSDEVLRKLVDDASKFLNDKVTKAVVTVPAYFN
DSQRTATKDAGRIAGLEVLRIINEPTAASLAYGFERKNNETILVFDLGGGTFDVSVLEVGDGVFEVLSTSGDTHLGGDDFDKRIVDWLAGDFKSNEGIDL
LKDKQALQRLTETAEKAKMELSSLTQANISLPFITATAEGPKHIETTLTRAKFEELCSDLLDRLKTPVENSLRDAKLTFKDLDEVILVGGSTRIPSVQGL
VKKLTGKDPNVTVNPDEVVALGAAVQAGVLSGDVSDIVLLDVTPLSIGLETLGGVMTKIIPRNTTLPSSKSEVFSTAADGQTSVEINVLQGEREFVRDNK
SLGSFRLDGIPPAPRGVPQIEVKFDIDANGILSVSAADKGTGKKQDITITGASTLPSDEVERMVQEAERFSKEDKEKRDAIDTKNQADSVVYQTEKQLKE
LGDKVPGEVKEKVESKLQELRDAIAGGSTQTMKDAMTTLNQEVMQLGQSLYNQQGSGAGEGPTTPGSESGPSSSSDSSSKGPDGDVIDADFTDSK
|
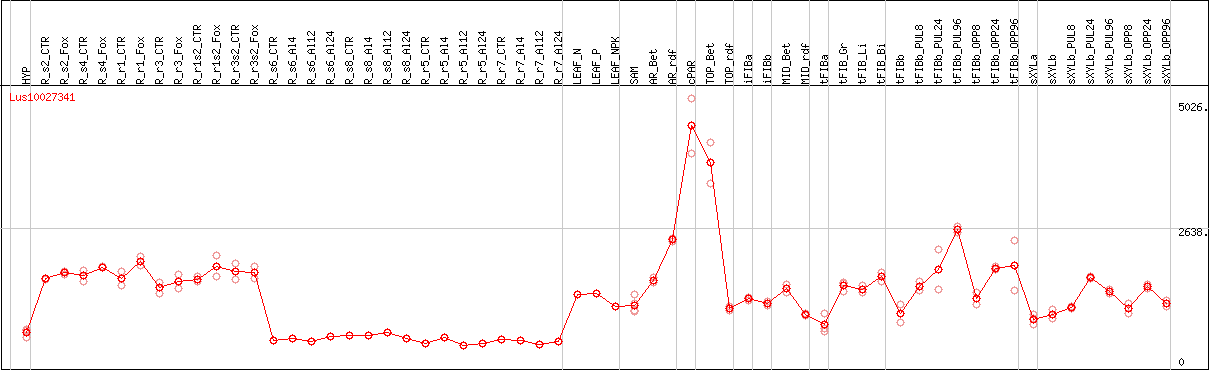
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027341 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
