External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G23730 246 / 1e-80
EFO2, RUP2
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G52250 218 / 2e-69
EFO1, RUP1
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 1, EARLY FLOWERING BY OVEREXPRESSION 1, Transducin/WD40 repeat-like superfamily protein (.1)
AT1G53090 154 / 1e-42
SPA4
SPA1-related 4 (.1.2)
AT3G15354 146 / 7e-40
SPA3
SPA1-related 3 (.1)
AT2G32950 143 / 6e-39
FUS1, EMB168, DET340, ATCOP1, COP1
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
AT2G46340 135 / 6e-36
SPA1
SUPPRESSOR OF PHYA-105 1, SPA (suppressor of phyA-105) protein family (.1)
AT4G11110 134 / 2e-35
SPA2
SPA1-related 2 (.1)
AT1G15440 52 / 1e-07
ATPWP2
\(PERIODIC TRYPTOPHAN PROTEIN 2, periodic tryptophan protein 2 (.1.2)
AT1G49040 48 / 3e-06
SCD1
STOMATAL CYTOKINESIS-DEFECTIVE 1, stomatal cytokinesis defective / SCD1 protein (SCD1) (.1), stomatal cytokinesis defective / SCD1 protein (SCD1) (.2), stomatal cytokinesis defective / SCD1 protein (SCD1) (.3)
AT1G29260 47 / 4e-06
PEX7, ATPEX7
ARABIDOPSIS PEROXIN 7, peroxin 7 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005742
352 / 4e-123
AT5G52250 226 / 3e-72
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 1, EARLY FLOWERING BY OVEREXPRESSION 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10035332
144 / 2e-39
AT2G32950 1088 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10029990
144 / 3e-39
AT2G32950 1088 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10043323
143 / 7e-39
AT3G15354 906 / 0.0
SPA1-related 3 (.1)
Lus10019475
142 / 2e-38
AT3G15354 992 / 0.0
SPA1-related 3 (.1)
Lus10033942
121 / 3e-31
AT4G11110 766 / 0.0
SPA1-related 2 (.1)
Lus10032358
91 / 2e-20
AT4G11110 784 / 0.0
SPA1-related 2 (.1)
Lus10040449
49 / 2e-06
AT1G49040 1816 / 0.0
STOMATAL CYTOKINESIS-DEFECTIVE 1, stomatal cytokinesis defective / SCD1 protein (SCD1) (.1), stomatal cytokinesis defective / SCD1 protein (SCD1) (.2), stomatal cytokinesis defective / SCD1 protein (SCD1) (.3)
Lus10023563
49 / 2e-06
AT1G49040 1743 / 0.0
STOMATAL CYTOKINESIS-DEFECTIVE 1, stomatal cytokinesis defective / SCD1 protein (SCD1) (.1), stomatal cytokinesis defective / SCD1 protein (SCD1) (.2), stomatal cytokinesis defective / SCD1 protein (SCD1) (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G140200
256 / 3e-84
AT5G23730 427 / 3e-149
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.015G143200
254 / 7e-84
AT5G23730 395 / 2e-137
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.011G119500
147 / 4e-40
AT3G15354 982 / 0.0
SPA1-related 3 (.1)
Potri.014G159300
145 / 6e-40
AT2G32950 1091 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.001G399600
140 / 1e-37
AT3G15354 1003 / 0.0
SPA1-related 3 (.1)
Potri.014G095300
139 / 3e-37
AT2G46340 846 / 0.0
SUPPRESSOR OF PHYA-105 1, SPA (suppressor of phyA-105) protein family (.1)
Potri.003G137100
133 / 3e-35
AT4G11110 1016 / 0.0
SPA1-related 2 (.1)
Potri.001G094500
132 / 4e-35
AT4G11110 1016 / 0.0
SPA1-related 2 (.1)
Potri.004G002700
131 / 6e-35
AT2G32950 832 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.011G021700
129 / 1e-34
AT2G32950 686 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
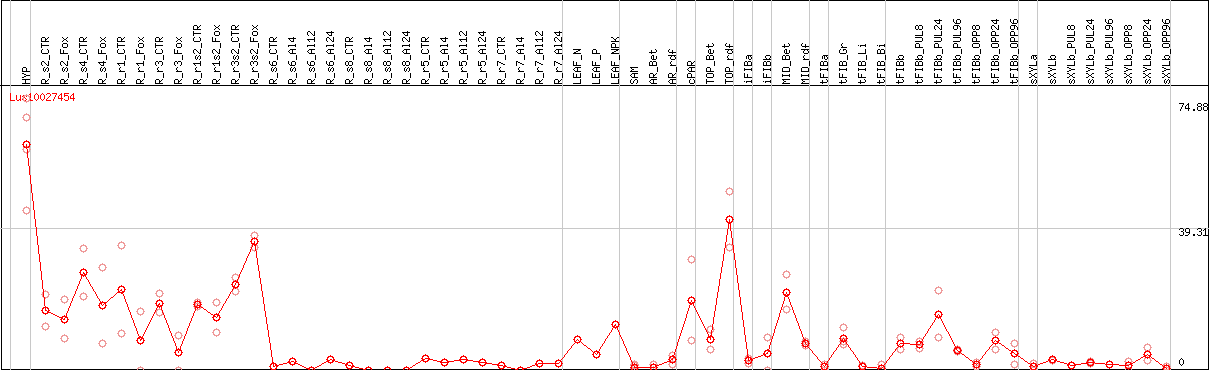
>Lus10027454 pacid=23144484 polypeptide=Lus10027454 locus=Lus10027454.g ID=Lus10027454.BGIv1.0 annot-version=v1.0
ATGGAGCGTCGACTATGGTTGTTTCGGCCTGTAGGCGCCAGCTGCGGTGTGGTCGGCGGCGGCGCCACGGCGGTTCTCTGCGCCTCGGGTTCCGACGACG
GGACTACCCACGTGTGGGACCCGCGATGCGAGGGAGGGGAATGCGTGGCTGTAATACGGCCAAGCGGGTCCGGATCCGCTGTATGCTGCGTGGAGTTCAA
TCGTTCTCGTGGGACGACGCTAGCCGTGGGATGCGCTGACCGGAAGGTGTACGTGTACGACTGCCGCATGCTGGTCCACCCTCTCCTTACCATGGGCGGG
CACACCAAAACGGTAACGTACGTCAAGTTCCTCGACGAAACGACTCTGGTCTCCGCGTCTATAGACGGGAGCTTGAAGCTGTGGGATCTGGGTAACCACT
CGGTCGAACCGGTTCGGACCTACCGCGGACATTCCAACGGTAGAAACTTCGTAGGGTTATCGGTTTGGAATGGGGGCGGGTTGCTGGGTTGCGGTTCGGA
GAGTAACCAGGCGTTCGTGTATGATCGGAGGTGGGGCGAACCGATTTGGATGCATGGTTTCAAACCGGTTGAACCAAGCACCGACGGGTTCGTGAGCAGT
GTGTGCTGGAAGCAAAGTGTGGACGACAATCTCGACAGCAGACAATGTACGTTCGTGGCCGGCGGATCGGATGGAACGTTGCGAGTTTTCGAAGGGATGA
AGAAGAAGATGACGGCGCCGGCTACTTCTTCCGTTGCGCCGACGAACACATGA
AA sequence
>Lus10027454 pacid=23144484 polypeptide=Lus10027454 locus=Lus10027454.g ID=Lus10027454.BGIv1.0 annot-version=v1.0
MERRLWLFRPVGASCGVVGGGATAVLCASGSDDGTTHVWDPRCEGGECVAVIRPSGSGSAVCCVEFNRSRGTTLAVGCADRKVYVYDCRMLVHPLLTMGG
HTKTVTYVKFLDETTLVSASIDGSLKLWDLGNHSVEPVRTYRGHSNGRNFVGLSVWNGGGLLGCGSESNQAFVYDRRWGEPIWMHGFKPVEPSTDGFVSS
VCWKQSVDDNLDSRQCTFVAGGSDGTLRVFEGMKKKMTAPATSSVAPTNT