External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G52790 298 / 1e-97
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
AT1G47330 260 / 9e-83
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
AT4G33700 249 / 1e-79
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
AT1G03270 245 / 3e-77
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
AT2G14520 242 / 3e-77
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
AT4G14240 229 / 3e-71
CBS domain-containing protein with a domain of unknown function (DUF21) (.1), CBS domain-containing protein with a domain of unknown function (DUF21) (.2)
AT4G14230 228 / 7e-71
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
AT3G13070 42 / 0.0007
CBS domain-containing protein / transporter associated domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027528
403 / 5e-139
AT5G52790 560 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10039289
396 / 3e-136
AT5G52790 557 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10039288
363 / 2e-123
AT5G52790 449 / 1e-154
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10005259
263 / 3e-85
AT2G14520 679 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10032715
260 / 4e-83
AT1G47330 646 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10030665
250 / 8e-82
AT2G14520 498 / 3e-178
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10012476
248 / 3e-79
AT2G14520 591 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10034024
243 / 1e-76
AT4G14230 675 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Lus10004429
239 / 7e-75
AT4G14230 654 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G147900
317 / 6e-105
AT5G52790 580 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Potri.001G288000
258 / 5e-83
AT2G14520 620 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Potri.014G012400
256 / 2e-82
AT2G14520 516 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Potri.014G196900
256 / 1e-80
AT1G47330 575 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Potri.009G082300
249 / 1e-79
AT2G14520 593 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Potri.008G202100
240 / 2e-75
AT4G14240 632 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1), CBS domain-containing protein with a domain of unknown function (DUF21) (.2)
Potri.010G030200
240 / 2e-75
AT4G14240 611 / 0.0
CBS domain-containing protein with a domain of unknown function (DUF21) (.1), CBS domain-containing protein with a domain of unknown function (DUF21) (.2)
Potri.002G260401
55 / 1e-09
AT1G47330 58 / 3e-11
CBS domain-containing protein with a domain of unknown function (DUF21) (.1)
Potri.001G362100
42 / 0.0004
AT1G55930 858 / 0.0
CBS domain-containing protein / transporter associated domain-containing protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00571
CBS
CBS domain
PF01595
DUF21
Cyclin M transmembrane N-terminal domain
Representative CDS sequence
>Lus10027527 pacid=23144517 polypeptide=Lus10027527 locus=Lus10027527.g ID=Lus10027527.BGIv1.0 annot-version=v1.0
ATGGTCCGTCCGAAACAAAATATTTCAAAGTTTGTTAAACTTACGTTCCTGCTCGCGGCACTTCCGATCTTCCTGGACTCAATTCTTCCTCCTTGGGCAG
CCATTCTCATATCAGTCACTCTCATTCTTGCCTTTGGTGAGATTATACCGCAATCAATTTGTTCAAGGTATGGACTCAGTGTTGGTGCAAAGCTGTCTCC
TATTGTTAAAGTCATTGTCATACTCCTTTTTCCAGTTGCTTACCCCATCAGTAAGCTGCTTGACTGGGTTCTTGGGAAGAAGCATTCGACACTCCTACGG
ACGAGCGGAGCTCAGAACCTTGGTGGACTTCCATGGAAACGAGGGAGAGGAGGAGAGTTGACACATGATGAGACAACAATCATAACTGGAGCTTTGGATA
TGACACAGAAGACTGCCAAAGATGCCATGACTCCCACTTCCAAAATCTTCTCTCTTGACATCAATTCTAAGCTTGATGAGAAAACAATGGAACTTATAAT
TAGCCATGGACATAGCCGTATACCCATCTACATGGGCAGCCCTAACAACATTGTTGGCCTCATTTTGGTTAAGAATTTGATCAAGTGTTGCTTTGAGGAT
GAAATACCGATTAGGGAACTTACAATCAGAAAGATTCCAAGAAAGGTCAGAGCCGCATGGCCATTGTCGTCAAGCAGAAGCACGATCCTAATAACAAAGT
CACCGAATTTCGAAAACCCGAAAGCAAAATCCGGAGTTTTAGAGATCAAAGTTGATCCCAATTCAAAACCAGGAGGCATAGAGAAGAAAGGAAATGGAGC
CTGCTTGAATATGATCCAAGATAAACCCATTATTGGTAAGAAAGATTTGGAGACGATCTCGTTCCCTGACATAGAGGTGATCGGCATCATAACAATGGAG
GATGTCATGGACAAACTTCTGCAGCAAGAGATTCTGGACGAGACCGACAAAACGACGACGTTTACAACCGGTAAGATCTGCAGAATTCAACTCTAA
AA sequence
>Lus10027527 pacid=23144517 polypeptide=Lus10027527 locus=Lus10027527.g ID=Lus10027527.BGIv1.0 annot-version=v1.0
MVRPKQNISKFVKLTFLLAALPIFLDSILPPWAAILISVTLILAFGEIIPQSICSRYGLSVGAKLSPIVKVIVILLFPVAYPISKLLDWVLGKKHSTLLR
TSGAQNLGGLPWKRGRGGELTHDETTIITGALDMTQKTAKDAMTPTSKIFSLDINSKLDEKTMELIISHGHSRIPIYMGSPNNIVGLILVKNLIKCCFED
EIPIRELTIRKIPRKVRAAWPLSSSRSTILITKSPNFENPKAKSGVLEIKVDPNSKPGGIEKKGNGACLNMIQDKPIIGKKDLETISFPDIEVIGIITME
DVMDKLLQQEILDETDKTTTFTTGKICRIQL
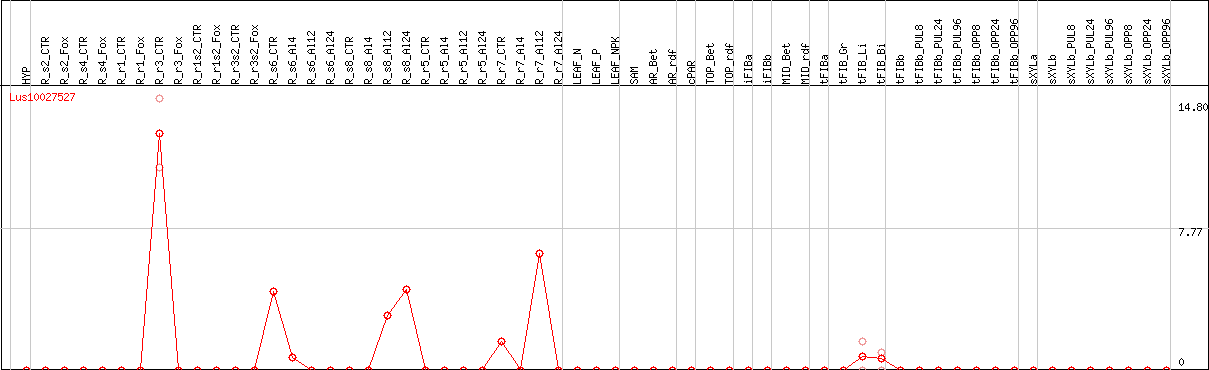
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027527 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.