External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G49142 205 / 1e-62
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G12770 130 / 4e-35
MEF22
mitochondrial editing factor 22 (.1)
AT4G25270 124 / 1e-33
OTP70
organelle transcript processing 70, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G33680 124 / 5e-33
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G15510 124 / 7e-33
VAC1, ATECB2
VANILLA CREAM 1, ARABIDOPSIS EARLY CHLOROPLAST BIOGENESIS2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G18520 121 / 3e-32
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G37170 120 / 2e-31
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G04840 119 / 2e-31
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G16470 118 / 2e-31
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G05240 118 / 5e-31
MEF19
mitochondrial editing factor 19 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039703
133 / 2e-36
AT4G18750 394 / 6e-127
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10018492
128 / 1e-34
AT4G18750 385 / 2e-123
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10031714
123 / 1e-32
AT4G25270 672 / 0.0
organelle transcript processing 70, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10004081
122 / 2e-32
AT3G63370 893 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 86, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10000706
122 / 4e-32
AT3G12770 878 / 0.0
mitochondrial editing factor 22 (.1)
Lus10024876
122 / 4e-32
AT3G12770 886 / 0.0
mitochondrial editing factor 22 (.1)
Lus10029436
120 / 9e-32
AT3G08820 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10026460
120 / 1e-31
AT1G15510 1030 / 0.0
VANILLA CREAM 1, ARABIDOPSIS EARLY CHLOROPLAST BIOGENESIS2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10001220
119 / 3e-31
AT3G57430 1110 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10027562 pacid=23144569 polypeptide=Lus10027562 locus=Lus10027562.g ID=Lus10027562.BGIv1.0 annot-version=v1.0
ATGGACGGAAATTACCGCTCAAACCCTTCCTTAGGGATCAAACTGATGAGGGCTTACGCAGCCTCTGGTCACGCCAGAGATGCACGCCGCATATTCGACG
GAATTACTGAGAAGAATGTCGTTTTCTTCAATGTCATGATTCGGAGCTACGTTAACAACCACTTCCATCGTGATGCGTTGGCTTTGTACAAAAACATATT
GAGTTCTGAAATCGTTCCTGATCATTACACGTTTCCTTGTGTTCTAAAGGCATGCTCTGGATCCGATAATCTGTGTCTCGGTGTCCAGATTCACGGTTGC
GTTATGAAAAATGGGCTCGACTCGAATCGGTACATAGTTAATTGCCTGGTTGCAATGTACGGGAAATGCAACCGTTTAAAGGAAGCTCGCCTGGTGCTCG
ATGAAATGCCTAGTAGAGATGTGGTTACGTGGAGTTCTTTGATTGCCGGGTGTGCTCAGAACGGACGTTTCGATGAGGCACTGGATTGTTGCAGAGAGAT
GGAGAGTTTGAAGCTTAAACCGGATTCGTCCACGATTCTCCCACAATGGCTAGCCCCCCACCGGCTGTGA
AA sequence
>Lus10027562 pacid=23144569 polypeptide=Lus10027562 locus=Lus10027562.g ID=Lus10027562.BGIv1.0 annot-version=v1.0
MDGNYRSNPSLGIKLMRAYAASGHARDARRIFDGITEKNVVFFNVMIRSYVNNHFHRDALALYKNILSSEIVPDHYTFPCVLKACSGSDNLCLGVQIHGC
VMKNGLDSNRYIVNCLVAMYGKCNRLKEARLVLDEMPSRDVVTWSSLIAGCAQNGRFDEALDCCREMESLKLKPDSSTILPQWLAPHRL
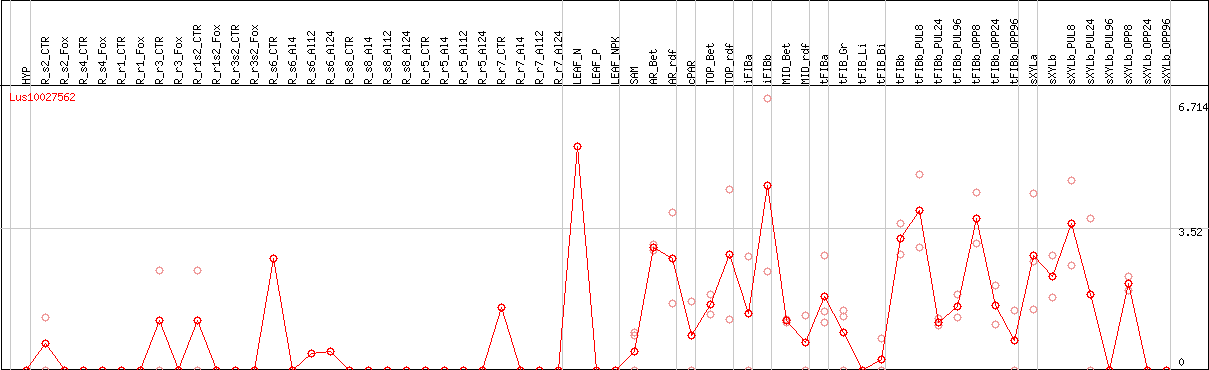
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027562 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.