External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G20200 154 / 8e-43
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT2G24370 146 / 4e-40
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT2G07020 143 / 3e-39
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT5G35380 139 / 9e-38
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT1G78940 136 / 1e-36
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1.2)
AT4G31230 134 / 7e-36
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT1G16760 131 / 5e-35
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT1G17540 119 / 7e-31
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT5G20310 114 / 1e-29
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT1G72760 105 / 5e-26
Protein kinase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008542
295 / 3e-95
AT2G24370 823 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10021596
214 / 7e-65
AT2G24370 828 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10040573
185 / 4e-54
AT2G24370 815 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10020184
144 / 3e-42
AT2G24370 286 / 2e-91
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10042205
146 / 3e-40
AT1G16760 691 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10008609
143 / 2e-39
AT1G16760 417 / 3e-137
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10026989
142 / 1e-38
AT2G24370 1014 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10024212
136 / 1e-36
AT2G24370 780 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Lus10017145
129 / 3e-35
AT5G35380 207 / 3e-60
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G001900
145 / 6e-40
AT1G78940 757 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1.2)
Potri.004G170901
144 / 7e-40
AT1G78940 584 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1.2)
Potri.004G170742
144 / 2e-39
AT1G78940 786 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1.2)
Potri.018G002400
135 / 2e-36
AT2G24370 956 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Potri.014G001700
130 / 2e-34
AT1G78940 783 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1.2)
Potri.018G061600
123 / 4e-32
AT5G12000 714 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Potri.006G078500
123 / 4e-32
AT2G24370 824 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Potri.001G198300
110 / 1e-27
AT5G12000 639 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Potri.006G225300
110 / 1e-27
AT5G12000 749 / 0.0
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Potri.018G147200
96 / 7e-23
AT5G20310 162 / 1e-45
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0039
HUP
PF00582
Usp
Universal stress protein family
Representative CDS sequence
>Lus10027592 pacid=23151254 polypeptide=Lus10027592 locus=Lus10027592.g ID=Lus10027592.BGIv1.0 annot-version=v1.0
ATGTCGAAGAAGATTGGAGAAACCAAGGAGTTCCAACAAGTGGTGGTGGCTGTAGACACAGACAAGGGAAGCCAGCATGCTCTGAAATGGGCAACTGATC
ATCTCCTCACCAAAGGCCAAACTGTTGTTCTCCTTCACGTCAAGCCCCGTCCATCTTCTTCTAATTCCTCTGTTTCAATGGTTGATGATGATCGTAACTC
ACAACCAAATCAAGCGCATGCCAGAGAACTGTTGCTTCCCTTCCGTTGCTTCTGCAGCCGCAAAGATTTTAAATGCAGTGAAGTTGTAGTCGAGGATTAC
GACATCGGCAAAGCAATCGTAGACTACGTGCATGCAAATTCTATCGATGCTTTGGTGCTCGGTTCTCCATCCAAAAGTGGCCTTGTCAGGAAGTTCAGGG
TGTCGGATGTTCCAACCAATGTAGCCAAACTGGCACCAGATTTCTGCACTGTTTACATCATCGGTAAAGGGAAGGTATCGTCGGTAAAATCTGCGACCGC
TTCGGTGCCTGCTAAACCTGTAGCCGCCATCCAAAGTTTCATGCCAGTGGCAGTCTCTGCATCATCAAGATCATCGTCCCCAAGCCGAAATTTCTCAAAT
CTCGAACCTTCTTCTACGCCGCACCAAAGAGGTTTGCTTTCCTTACCAGCCTATGAGTTCTTTTACTTCCTGCCAGACGTTATAGCATAA
AA sequence
>Lus10027592 pacid=23151254 polypeptide=Lus10027592 locus=Lus10027592.g ID=Lus10027592.BGIv1.0 annot-version=v1.0
MSKKIGETKEFQQVVVAVDTDKGSQHALKWATDHLLTKGQTVVLLHVKPRPSSSNSSVSMVDDDRNSQPNQAHARELLLPFRCFCSRKDFKCSEVVVEDY
DIGKAIVDYVHANSIDALVLGSPSKSGLVRKFRVSDVPTNVAKLAPDFCTVYIIGKGKVSSVKSATASVPAKPVAAIQSFMPVAVSASSRSSSPSRNFSN
LEPSSTPHQRGLLSLPAYEFFYFLPDVIA
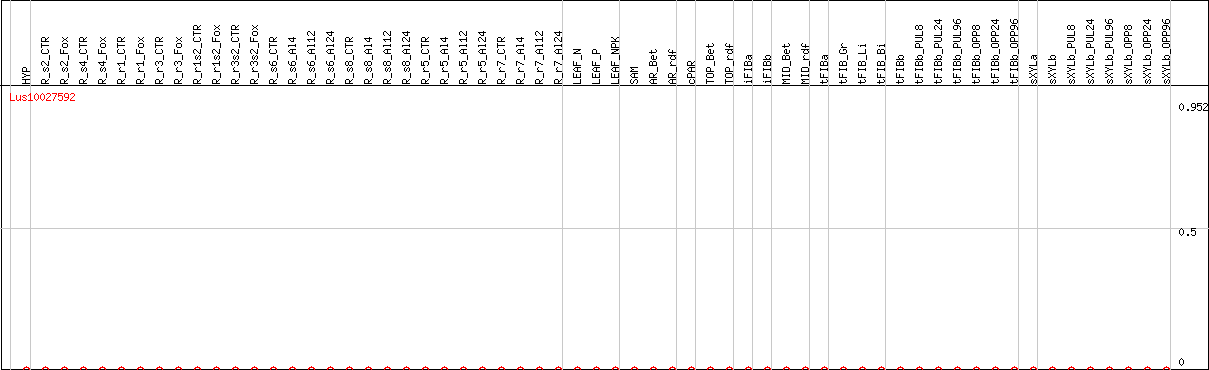
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027592 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.