External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06839 208 / 1e-66
bZIP
TGA10, bZIP65
TGACG \(TGA\) motif-binding protein 10, bZIP transcription factor family protein (.1.2.3)
AT5G06960 175 / 8e-55
bZIP
TGA5, OBF5
TGACG MOTIF-BINDING FACTOR 5, OCS-element binding factor 5 (.1.2)
AT3G12250 172 / 8e-54
bZIP
BZIP45, TGA6
TGACG motif-binding factor 6 (.1.2.3.4.5)
AT1G68640 176 / 9e-54
bZIP
PAN
PERIANTHIA, bZIP transcription factor family protein (.1)
AT5G06950 172 / 1e-53
bZIP
TGA2, AHBP-1B
bZIP transcription factor family protein (.1.2.3.4)
AT1G08320 164 / 4e-50
bZIP
TGA9, bZIP21
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
AT1G77920 150 / 6e-45
bZIP
bZIP transcription factor family protein (.1)
AT1G22070 141 / 5e-41
bZIP
TGA3
TGA1A-related gene 3 (.1)
AT5G10030 140 / 5e-41
bZIP
OBF4, TGA4
OCS ELEMENT BINDING FACTOR 4, TGACG motif-binding factor 4 (.1.2)
AT5G65210 132 / 1e-38
bZIP
TGA1
bZIP transcription factor family protein (.1.2.3.4.5.6)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035499
276 / 4e-94
AT5G06839 377 / 1e-129
TGACG \(TGA\) motif-binding protein 10, bZIP transcription factor family protein (.1.2.3)
Lus10023822
241 / 2e-78
AT5G06839 491 / 2e-171
TGACG \(TGA\) motif-binding protein 10, bZIP transcription factor family protein (.1.2.3)
Lus10026166
192 / 3e-61
AT3G12250 514 / 0.0
TGACG motif-binding factor 6 (.1.2.3.4.5)
Lus10024140
189 / 3e-60
AT3G12250 532 / 0.0
TGACG motif-binding factor 6 (.1.2.3.4.5)
Lus10008658
184 / 3e-58
AT3G12250 521 / 0.0
TGACG motif-binding factor 6 (.1.2.3.4.5)
Lus10020610
173 / 5e-52
AT1G08320 633 / 0.0
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
Lus10004876
172 / 2e-51
AT1G08320 635 / 0.0
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
Lus10041475
170 / 3e-51
AT1G68640 444 / 4e-153
PERIANTHIA, bZIP transcription factor family protein (.1)
Lus10043387
162 / 2e-48
AT1G08320 481 / 7e-168
TGACG \(TGA\) motif-binding protein 9, bZIP transcription factor family protein (.1.2.3)
PFAM info
Representative CDS sequence
>Lus10027802 pacid=23156978 polypeptide=Lus10027802 locus=Lus10027802.g ID=Lus10027802.BGIv1.0 annot-version=v1.0
ATGTACGTGGACAACTGCTTGGCCCACTACGACGTCGTTATGACTCTCAAAGGGATGGCGGCCAAAGCGGACGTCTTCCATTTGGTGTCCGGAATGTGGA
AGACGCCAGCCGAGCGTTGCTTCATTTGGATCGGCGGCTTCCGTCCTTCCGATCTCATCAAGGTAATAGTGAGCCAGATAGAGCCACTAACGGAGCAGCA
AATCCTTGGGGTATGCGGATTGCAGCAGTCGACCCAGGAAGCCGAGGAAGCCCTATCTCAAGGGCTTGAAGCGCTCAACCAGTCGATTTCAGAGACGATA
GCTTCAGACTCGTTGATCAACTCCCCTCCAAAGAACATGGCCAACTACATGGGTCAGATGGCCTTGGCCATCAACAAGCTCTCTGCCATCGAAGGCTTCG
TACGACAGAATATTGAGAGGACTCATCTCATTAATTTGAATTGTTGGGAGAGGTACATAGGTTCACAATGA
AA sequence
>Lus10027802 pacid=23156978 polypeptide=Lus10027802 locus=Lus10027802.g ID=Lus10027802.BGIv1.0 annot-version=v1.0
MYVDNCLAHYDVVMTLKGMAAKADVFHLVSGMWKTPAERCFIWIGGFRPSDLIKVIVSQIEPLTEQQILGVCGLQQSTQEAEEALSQGLEALNQSISETI
ASDSLINSPPKNMANYMGQMALAINKLSAIEGFVRQNIERTHLINLNCWERYIGSQ
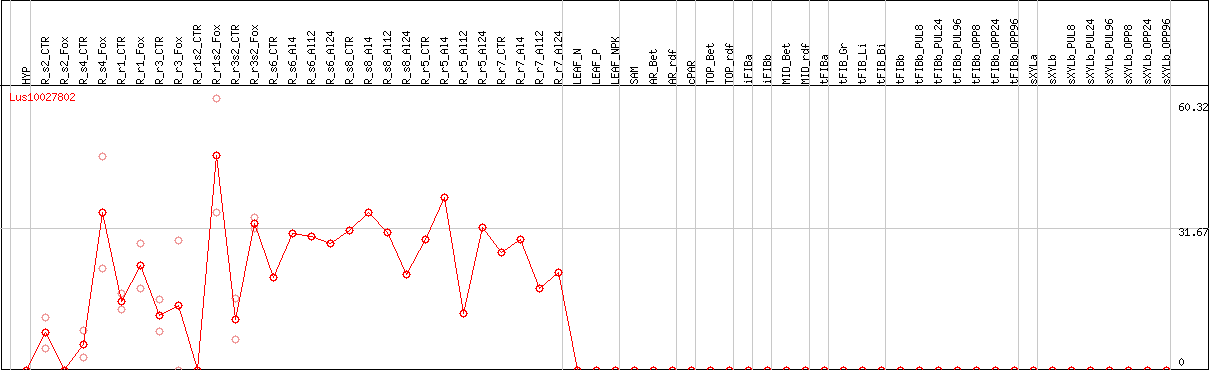
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027802 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.