External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G01340 526 / 0
AtmSFC1
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
AT3G54110 114 / 2e-29
ATUCP1, UCP1, UCP2, UCP, ATPUMP1
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
AT1G14140 110 / 8e-28
Mitochondrial substrate carrier family protein (.1)
AT5G58970 105 / 2e-26
ATUCP2
uncoupling protein 2 (.1.2)
AT2G37890 105 / 5e-26
Mitochondrial substrate carrier family protein (.1)
AT1G34065 102 / 1e-24
SAMC2
S-adenosylmethionine carrier 2 (.1)
AT4G03115 101 / 1e-24
Mitochondrial substrate carrier family protein (.1)
AT4G24570 99 / 9e-24
DIC2
dicarboxylate carrier 2 (.1)
AT3G53940 99 / 2e-23
Mitochondrial substrate carrier family protein (.1)
AT5G19760 98 / 2e-23
Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005045
633 / 0
AT5G01340 518 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Lus10035538
108 / 3e-27
AT3G54110 512 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10027755
106 / 1e-26
AT3G54110 515 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10007883
107 / 4e-26
AT5G19760 536 / 0.0
Mitochondrial substrate carrier family protein (.1)
Lus10017187
105 / 6e-26
AT3G54110 501 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10021123
103 / 2e-25
AT3G54110 521 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10040709
102 / 7e-25
AT5G58970 512 / 0.0
uncoupling protein 2 (.1.2)
Lus10034104
101 / 2e-24
AT1G25380 414 / 3e-145
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Lus10000906
100 / 4e-24
AT5G46800 482 / 1e-173
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G119500
548 / 0
AT5G01340 542 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Potri.016G110100
108 / 3e-27
AT3G54110 483 / 5e-174
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Potri.006G095500
107 / 7e-27
AT3G54110 513 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Potri.001G247800
107 / 1e-26
AT5G58970 456 / 3e-163
uncoupling protein 2 (.1.2)
Potri.010G121500
105 / 1e-25
AT1G25380 395 / 2e-137
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Potri.016G119650
97 / 2e-25
AT5G01340 88 / 5e-23
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Potri.003G053900
102 / 5e-25
AT1G79900 419 / 2e-148
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Potri.001G182000
102 / 6e-25
AT1G79900 424 / 1e-150
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Potri.007G019000
100 / 2e-24
AT5G66380 463 / 8e-166
folate transporter 1 (.1)
Potri.002G200900
99 / 1e-23
AT2G47490 471 / 1e-168
NAD+ transporter 1, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 1, NAD+ transporter 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Lus10027817 pacid=23156881 polypeptide=Lus10027817 locus=Lus10027817.g ID=Lus10027817.BGIv1.0 annot-version=v1.0
ATGGCAGAGCCCAACAACCAAACCCCGACCAACCGCCATCCGAACAAGGCCACAATCCCGCCGTACATGAAGGCGATGTCGGGATCGCTCGGAGGGATAA
TGGAGGCCTGCTGTCTACAGCCGATAGACGTCATCAAGACTCGCCTGCAGCTCGACCGGAGCGGGGCTTACAGGGGAATCGTCCACTGTGGATCCACGGT
TTGCCAGACGGAAGGGGTTCGAGCTCTGTGGAAAGGATTGACCCCTTTCGCCACTCACTTGACTTTGAAATACTCGCTCCGGATGGGGACCAACGCCGTG
CTTCAGTCCGCGTTCAAGGATTCGCAGACTGGACACCTCAGCAATGGCTCCAGGCTCATGTCTGGTTTTGGTGCTGGCGTTTTGGAGGCTCTCGTCATCG
TTACTCCTTTCGAGGTGGTGAAGATCAGGCTACAGCAGCAAAAGGGATTAAGTTCTGATCTTCTCAAGTACAAAGGACCAGTACATTGTGCCCGTTTGAT
CGTCCGAGAGGAAGGAATCCTAGGTCTGTGGGCAGGTGCGGCACCAACCGTGATGCGAAATGGCACGAACCAGGCAGCCATGTTCACAGCCAAGAACGCA
TTCGACATCCTATTGTGGAAGAAGAACGAAGGGGACGGGAAAGTTCTCCTACCGTGGCAGTCCATGATCTCAGGATTTCTCGCAGGAACAGCCGGTCCGA
TCTGCACAGGACCTTTCGACGTGGTGAAAACTCGGCTCATGGCTCAGAGTCGAGCAGGTGGTAATATGAAGTACAACGGAATGGTTCATGCGATCAGGAC
AATCTATGCAGAAGAAGGACTGCTTGCCTTGTGGAAAGGATTGATACCTCGGCTGATGAGGATCCCGCCCGGGCAGGCTATAATGTGGGCTGTGGCTGAC
CAGGTCATGGGTTTGTATGAGAAGAGGTATATACCCAGTGCAGCTTTGTAG
AA sequence
>Lus10027817 pacid=23156881 polypeptide=Lus10027817 locus=Lus10027817.g ID=Lus10027817.BGIv1.0 annot-version=v1.0
MAEPNNQTPTNRHPNKATIPPYMKAMSGSLGGIMEACCLQPIDVIKTRLQLDRSGAYRGIVHCGSTVCQTEGVRALWKGLTPFATHLTLKYSLRMGTNAV
LQSAFKDSQTGHLSNGSRLMSGFGAGVLEALVIVTPFEVVKIRLQQQKGLSSDLLKYKGPVHCARLIVREEGILGLWAGAAPTVMRNGTNQAAMFTAKNA
FDILLWKKNEGDGKVLLPWQSMISGFLAGTAGPICTGPFDVVKTRLMAQSRAGGNMKYNGMVHAIRTIYAEEGLLALWKGLIPRLMRIPPGQAIMWAVAD
QVMGLYEKRYIPSAAL
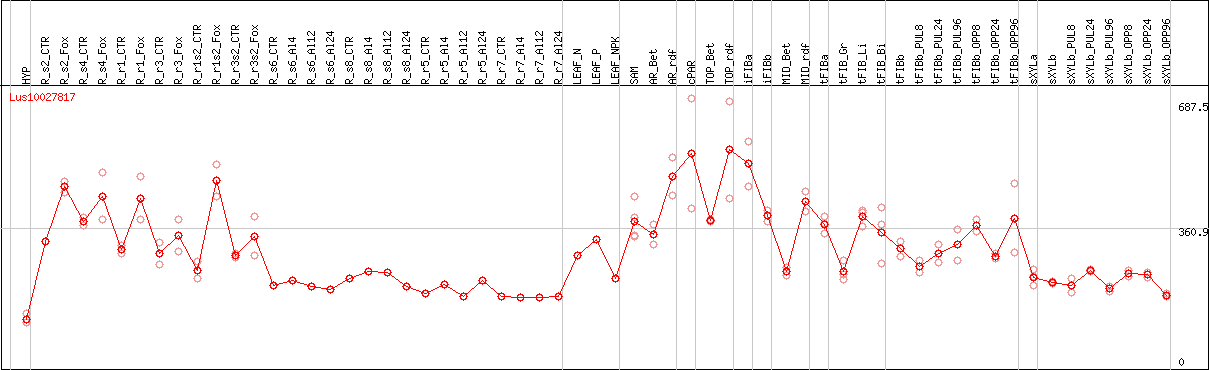
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027817 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.