External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G60170 197 / 7e-60
EMB1220
embryo defective 1220, pre-mRNA processing ribonucleoprotein binding region-containing protein (.1)
AT1G70400 141 / 1e-41
unknown protein
AT5G27120 83 / 1e-17
NOP56-like pre RNA processing ribonucleoprotein (.1)
AT3G05060 81 / 4e-17
NOP56-like pre RNA processing ribonucleoprotein (.1)
AT5G27140 77 / 8e-16
NOP56-like pre RNA processing ribonucleoprotein (.1)
AT1G56110 69 / 6e-13
NOP56
homolog of nucleolar protein NOP56 (.1)
AT3G12860 65 / 1e-11
NOP56-like pre RNA processing ribonucleoprotein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029168
234 / 1e-73
AT1G60170 707 / 0.0
embryo defective 1220, pre-mRNA processing ribonucleoprotein binding region-containing protein (.1)
Lus10012997
235 / 3e-73
AT1G60170 705 / 0.0
embryo defective 1220, pre-mRNA processing ribonucleoprotein binding region-containing protein (.1)
Lus10002830
135 / 1e-39
AT1G60170 135 / 7e-39
embryo defective 1220, pre-mRNA processing ribonucleoprotein binding region-containing protein (.1)
Lus10004143
85 / 2e-18
AT5G27120 730 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Lus10001716
84 / 7e-18
AT5G27120 722 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Lus10033846
82 / 2e-17
AT5G27120 758 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Lus10018995
82 / 2e-17
AT5G27120 734 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Lus10017214
60 / 5e-10
AT1G56110 783 / 0.0
homolog of nucleolar protein NOP56 (.1)
Lus10021096
60 / 7e-10
AT1G56110 782 / 0.0
homolog of nucleolar protein NOP56 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G176000
241 / 1e-76
AT1G60170 591 / 0.0
embryo defective 1220, pre-mRNA processing ribonucleoprotein binding region-containing protein (.1)
Potri.014G102800
239 / 5e-76
AT1G60170 610 / 0.0
embryo defective 1220, pre-mRNA processing ribonucleoprotein binding region-containing protein (.1)
Potri.013G032300
84 / 5e-18
AT3G05060 737 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Potri.005G045600
82 / 2e-17
AT5G27120 768 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Potri.015G092900
70 / 3e-13
AT3G12860 761 / 0.0
NOP56-like pre RNA processing ribonucleoprotein (.1)
Potri.012G095400
69 / 5e-13
AT1G56110 761 / 0.0
homolog of nucleolar protein NOP56 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01798
Nop
snoRNA binding domain, fibrillarin
Representative CDS sequence
>Lus10027881 pacid=23157082 polypeptide=Lus10027881 locus=Lus10027881.g ID=Lus10027881.BGIv1.0 annot-version=v1.0
ATGGCGATGAGGAAGGTGATGGATAATGAAGAAGGAGTGGCTATGAGGAGGAAGGCTAAGGAGCTTGGAAGAATGGCGAGGAGGGCGGTTGAGGACAGTG
GATCTTCGAATTCTAGTTTGAGCTGGGGCTCCTTCCCGAACTTCAAAATTTCCCGGCTCCGTTCACAGCAACAGCTTCTCATCCTTCGTCGGCCGGTTGA
GGACATCAGTAGCGGATTCAGGGGCCTCCCCAGGCGCCCGGACCCGTACATAGAGAACGAGCTTACCATCATCCACAACTTCATCCGCGCCAAGTACCGG
TCCAAGTTCCCCGAGCTGGAGTCTCTCGTGCGCGATCACATTGAATACGCTCGCGTGGTGAAGAGAATCGGGAACGAGATAGATATAATCCTCGTTGATC
TCGAAGGTCTTTTGCCATCAATCACCATTATGGTCGTTTCAGTGACTGCATTGACAACAAGCGGTAAACCTCTCCCCCTTGACGTTCTCGAAACGGTTAT
TGATGCTTGTGATCGGATTCTAACTCTGGATTCCGCAAAGAAGAAGGTTATCGATTTTGTGGAGAGCAGAATGGGACGCATCCCGAACTTGTCTACCATC
GTCTGGAGTTCCGTGGCAGCAAAGCTCATTTGCATTGCCGGTGGTCTCACTGCTTTAGCCAACATGCCTTCTTCCAATGTTGAACTTCTCGGAGCCAAGA
AGAAGAGTGCAACATCATCTCAATTTCGCTTTGGTTATATTGAGCAAACGGATGGCTGGCTTCGAAATCGACACTTGCAGCCTGAGTCGATGCTGAAAGT
TCAAGTCCTTGTGGAAGCCATGGAAGAACTCTTCGAGAAGAGATTGTTAAGAAGATAG
AA sequence
>Lus10027881 pacid=23157082 polypeptide=Lus10027881 locus=Lus10027881.g ID=Lus10027881.BGIv1.0 annot-version=v1.0
MAMRKVMDNEEGVAMRRKAKELGRMARRAVEDSGSSNSSLSWGSFPNFKISRLRSQQQLLILRRPVEDISSGFRGLPRRPDPYIENELTIIHNFIRAKYR
SKFPELESLVRDHIEYARVVKRIGNEIDIILVDLEGLLPSITIMVVSVTALTTSGKPLPLDVLETVIDACDRILTLDSAKKKVIDFVESRMGRIPNLSTI
VWSSVAAKLICIAGGLTALANMPSSNVELLGAKKKSATSSQFRFGYIEQTDGWLRNRHLQPESMLKVQVLVEAMEELFEKRLLRR
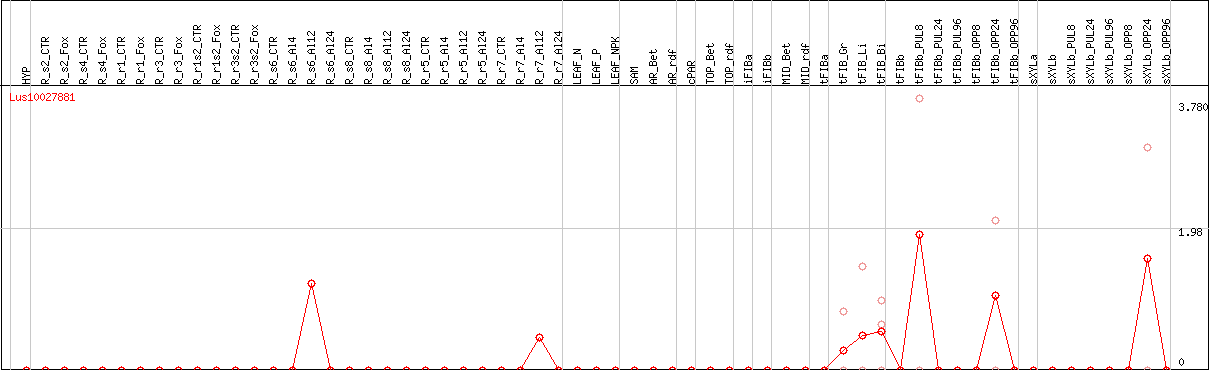
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027881 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.