External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G33985 110 / 3e-31
Protein of unknown function (DUF1685) (.1)
AT3G50350 99 / 1e-26
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
AT2G15590 85 / 3e-21
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
AT2G15610 65 / 2e-13
Protein of unknown function (DUF1685) (.1)
AT1G08790 61 / 6e-12
Protein of unknown function (DUF1685) (.1)
AT3G04700 59 / 3e-11
Protein of unknown function (DUF1685) (.1)
AT3G04710 58 / 3e-10
TPR10
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
AT5G28690 54 / 3e-09
Protein of unknown function (DUF1685) (.1)
AT2G31560 50 / 5e-08
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2)
AT2G43340 42 / 8e-05
Protein of unknown function (DUF1685) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002400
247 / 6e-85
AT4G33985 150 / 7e-47
Protein of unknown function (DUF1685) (.1)
Lus10014064
119 / 1e-34
AT4G33985 145 / 2e-45
Protein of unknown function (DUF1685) (.1)
Lus10019852
116 / 2e-33
AT4G33985 149 / 9e-47
Protein of unknown function (DUF1685) (.1)
Lus10021997
67 / 1e-14
AT1G08790 129 / 4e-39
Protein of unknown function (DUF1685) (.1)
Lus10042536
67 / 2e-14
AT1G08790 141 / 2e-43
Protein of unknown function (DUF1685) (.1)
Lus10006560
66 / 1e-13
AT1G08790 166 / 2e-52
Protein of unknown function (DUF1685) (.1)
Lus10005536
46 / 9e-07
AT5G28690 119 / 3e-35
Protein of unknown function (DUF1685) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G101300
116 / 1e-33
AT4G33985 130 / 2e-39
Protein of unknown function (DUF1685) (.1)
Potri.005G138100
96 / 4e-25
AT4G33985 139 / 2e-42
Protein of unknown function (DUF1685) (.1)
Potri.007G043900
88 / 3e-22
AT4G33985 134 / 3e-40
Protein of unknown function (DUF1685) (.1)
Potri.013G042400
67 / 4e-14
AT1G08790 151 / 2e-46
Protein of unknown function (DUF1685) (.1)
Potri.005G055100
62 / 5e-12
AT1G08790 126 / 1e-36
Protein of unknown function (DUF1685) (.1)
Potri.007G126700
51 / 3e-08
AT1G05870 188 / 7e-61
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2), Protein of unknown function (DUF1685) (.3), Protein of unknown function (DUF1685) (.4)
Potri.017G032800
47 / 6e-07
AT1G05870 190 / 1e-61
Protein of unknown function (DUF1685) (.1), Protein of unknown function (DUF1685) (.2), Protein of unknown function (DUF1685) (.3), Protein of unknown function (DUF1685) (.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07939
DUF1685
Protein of unknown function (DUF1685)
Representative CDS sequence
>Lus10027898 pacid=23156933 polypeptide=Lus10027898 locus=Lus10027898.g ID=Lus10027898.BGIv1.0 annot-version=v1.0
ATGGCGACCACCGAACAATCAAACCAAGCCGTACCGATGACAACTCGCCGCCGGCAGCAGAAGCCGGTTCTCCGGAAGATGAACTCATGGTCGCCTGACA
GCGGGAAGGAGGACGTGTGGCGCCGCCGCAAGGGGAACCACTGCTCCTCCTGCCGCCACCTCCACCTCCGCCGTGCCAATAGCAACGTCACCGACGAGGA
TCTCGACGAGCTCAAGGCCTGCATAGAGCTCGGGTTCGGATTCGACCCGGACTCCCGGCCGGATCTGGATCCCAGGCTCTCTGACGCCCTGCCGGCGCTC
GGACTGTACTGTGCTGTCAACAAGCAGTATAACAGAATCCTGTCGCGGTCAGCGTCGTCGGTTTCGTCAGACGGCGGCGCCGTAACTCCTCCTCCTCTCC
TCCCGTCGTCGTGGACCCCAGTAAGTGAAGATCCGGAGACGGTGAAGATGAAGCTGAGGCAGTGGGCGAGAGTGGTGGCGTGTTCAGTGAAGCAATTAGC
TGGGAATTAA
AA sequence
>Lus10027898 pacid=23156933 polypeptide=Lus10027898 locus=Lus10027898.g ID=Lus10027898.BGIv1.0 annot-version=v1.0
MATTEQSNQAVPMTTRRRQQKPVLRKMNSWSPDSGKEDVWRRRKGNHCSSCRHLHLRRANSNVTDEDLDELKACIELGFGFDPDSRPDLDPRLSDALPAL
GLYCAVNKQYNRILSRSASSVSSDGGAVTPPPLLPSSWTPVSEDPETVKMKLRQWARVVACSVKQLAGN
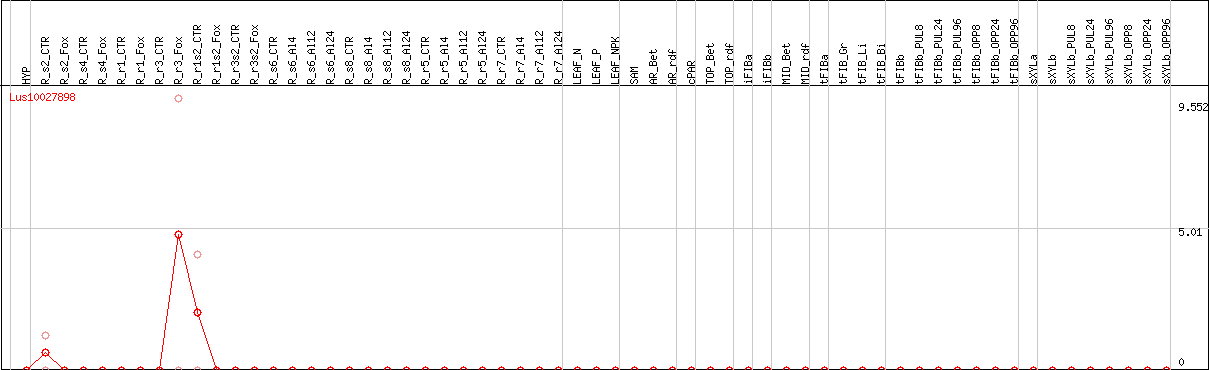
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027898 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.