External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G18990 69 / 2e-13
B3
REM39, VRN1
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
AT4G33280 59 / 5e-10
B3
REM16
AP2/B3-like transcriptional factor family protein (.1)
AT4G01580 56 / 4e-09
B3
AP2/B3-like transcriptional factor family protein (.1)
AT3G18960 56 / 5e-09
B3
AP2/B3-like transcriptional factor family protein (.1)
AT1G49480 47 / 3e-06
B3
REM19, RTV1
related to vernalization1 1 (.1)
AT1G49475 43 / 9e-05
B3
AP2/B3-like transcriptional factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012039
382 / 6e-135
AT3G18990 74 / 4e-15
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10027937
352 / 7e-123
AT3G18990 62 / 9e-11
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10012045
283 / 2e-96
ND 44 / 2e-05
Lus10010759
105 / 2e-26
AT3G18990 91 / 1e-20
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10010756
105 / 3e-26
AT3G18990 99 / 2e-23
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10035637
101 / 5e-24
AT3G18990 91 / 9e-20
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10025507
85 / 4e-19
AT3G18990 91 / 1e-21
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10041961
81 / 1e-17
AT3G18990 94 / 3e-22
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Lus10023079
79 / 2e-17
AT3G18990 69 / 3e-13
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G191500
74 / 8e-15
AT3G18990 189 / 5e-57
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.004G230100
63 / 5e-11
AT3G18990 161 / 1e-44
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.004G147200
61 / 3e-10
AT1G49480 110 / 5e-28
related to vernalization1 1 (.1)
Potri.014G057101
56 / 6e-09
AT3G18990 76 / 5e-16
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.007G035700
54 / 2e-08
AT4G01580 71 / 4e-15
AP2/B3-like transcriptional factor family protein (.1)
Potri.009G108500
50 / 9e-07
AT3G18990 395 / 2e-137
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.004G147000
49 / 1e-06
AT3G18990 258 / 3e-83
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.004G146900
48 / 4e-06
AT3G18990 353 / 6e-121
REDUCED VERNALIZATION RESPONSE 1, REPRODUCTIVE MERISTEM 39, AP2/B3-like transcriptional factor family protein (.1)
Potri.007G035500
41 / 0.0005
AT5G66980 102 / 4e-25
AP2/B3-like transcriptional factor family protein (.1)
Potri.014G036900
41 / 0.0006
AT4G33280 256 / 5e-83
AP2/B3-like transcriptional factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0405
DNA_b-psBarrel
PF02362
B3
B3 DNA binding domain
Representative CDS sequence
>Lus10027942 pacid=23156919 polypeptide=Lus10027942 locus=Lus10027942.g ID=Lus10027942.BGIv1.0 annot-version=v1.0
ATGTCTGCAGCCAACAAATTCCCATTCCTGCAATTCTTCAAGGTGATCTTGCCTGAAACCATTCATCACAAGGAAATGATGCTTCCTCGGAGGTTTATGA
AGAATTTCAAGGCCGAATTCTCTGGTTCCGAGCTGACATTGGAAGTTGCTGATGGAAAAACTTGGACAGTTGGATTGGAGGAAGAGGAAGGACATCTTAA
CCGAGTCTGGCTCGTCTCAGGATTCGAAGATTTCCTCAGGTTCTACTCCATCTCTCCGAAACAGATCCTCCTCTTCAGGTACCTCAAGCCGTCCCGTCTC
AAGGTCTTCATCTTGCACTCAACATCAATGGAGATAACTTATCCACCACTAGCCCCAGCCCCCGATTCCGACGAACTGTTGGACCGGCTTAAGAAGATGA
AGAAGATGCAACTGAGCAACAAGTTCAACTTAGCATTCGAGACGATGTCTACGAACAGCAAGAGGGCGGTCGAGAGAGCTCTGGAGGAGATGAATCCGGC
CAACATTTCTACACTGGTTGTCATAGCGAGTTATCACATACGCCCAGGAAACATGTATCTTCCAGTGAACTTCACTAAGAAGTTGTGCTCGAGGGATGAT
TCAACTACGGCAAGGCTGAAGCTCTATGAGGGGACGGATCAGGTGGTGGGCGAGTGGCCGATTGAATTCAACAAACTTCGAAGTCAGAGGAATCAGAAAA
ATGTCACAGGATGGAACAAGTTTCGCCGGGAAAACGAGTTGGTCGAGGGAGACGTTTGTCTTTTCGAGCTGATTCCTCGTTCCGAGGGGGAGATCTTACT
ACTCAAAGTTCGTATCGTCAGGGCTGCTGCTGAGTGA
AA sequence
>Lus10027942 pacid=23156919 polypeptide=Lus10027942 locus=Lus10027942.g ID=Lus10027942.BGIv1.0 annot-version=v1.0
MSAANKFPFLQFFKVILPETIHHKEMMLPRRFMKNFKAEFSGSELTLEVADGKTWTVGLEEEEGHLNRVWLVSGFEDFLRFYSISPKQILLFRYLKPSRL
KVFILHSTSMEITYPPLAPAPDSDELLDRLKKMKKMQLSNKFNLAFETMSTNSKRAVERALEEMNPANISTLVVIASYHIRPGNMYLPVNFTKKLCSRDD
STTARLKLYEGTDQVVGEWPIEFNKLRSQRNQKNVTGWNKFRRENELVEGDVCLFELIPRSEGEILLLKVRIVRAAAE
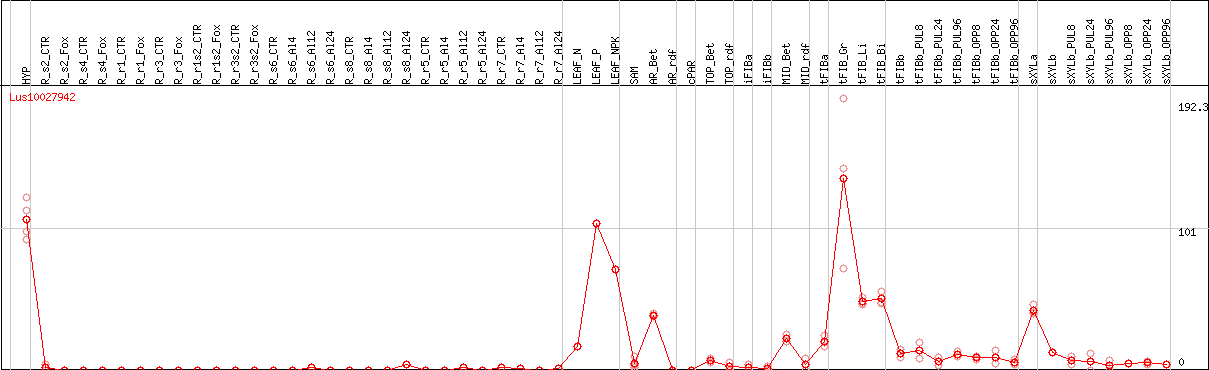
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027942 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.