External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G03440 161 / 1e-49
Leucine-rich repeat (LRR) family protein (.1)
AT4G03010 159 / 5e-49
RNI-like superfamily protein (.1)
AT3G17640 69 / 3e-15
Leucine-rich repeat (LRR) family protein (.1)
AT2G23950 44 / 2e-06
Leucine-rich repeat protein kinase family protein (.1)
AT1G13910 42 / 1e-05
Leucine-rich repeat (LRR) family protein (.1)
AT5G61240 41 / 2e-05
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
AT4G30520 40 / 5e-05
SARK
SENESCENCE-ASSOCIATED RECEPTOR-LIKE KINASE, Leucine-rich repeat protein kinase family protein (.1)
AT3G59510 38 / 0.0004
Leucine-rich repeat (LRR) family protein (.1)
AT1G50610 37 / 0.0008
Leucine-rich repeat protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008190
197 / 8e-64
AT1G03440 576 / 0.0
Leucine-rich repeat (LRR) family protein (.1)
Lus10042681
166 / 7e-53
AT4G03010 205 / 1e-63
RNI-like superfamily protein (.1)
Lus10029622
167 / 5e-52
AT1G03440 534 / 0.0
Leucine-rich repeat (LRR) family protein (.1)
Lus10031601
151 / 6e-46
AT1G03440 463 / 2e-163
Leucine-rich repeat (LRR) family protein (.1)
Lus10037108
45 / 1e-06
AT5G61240 550 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10026651
43 / 6e-06
AT5G61240 549 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10004662
43 / 7e-06
AT5G61240 543 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10036961
43 / 8e-06
AT5G61240 488 / 1e-175
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10015484
40 / 5e-05
AT2G23950 910 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G136100
170 / 4e-53
AT1G03440 541 / 0.0
Leucine-rich repeat (LRR) family protein (.1)
Potri.004G001000
82 / 1e-19
AT3G17640 386 / 1e-132
Leucine-rich repeat (LRR) family protein (.1)
Potri.011G023500
80 / 5e-19
AT3G17640 350 / 2e-118
Leucine-rich repeat (LRR) family protein (.1)
Potri.008G093400
45 / 1e-06
AT5G61240 513 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.010G160700
44 / 2e-06
AT5G61240 517 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.018G002600
39 / 0.0001
AT3G20190 540 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Potri.007G078100
38 / 0.0003
AT4G39400 1426 / 0.0
DWARF 2, CABBAGE 2, BRASSINOSTEROID INSENSITIVE 1, BR INSENSITIVE 1, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.005G037400
38 / 0.0003
AT3G47570 622 / 0.0
Leucine-rich repeat protein kinase family protein (.1)
Potri.005G086500
38 / 0.0003
AT4G39400 1398 / 0.0
DWARF 2, CABBAGE 2, BRASSINOSTEROID INSENSITIVE 1, BR INSENSITIVE 1, Leucine-rich receptor-like protein kinase family protein (.1)
Potri.019G078400
38 / 0.0005
AT1G72180 1060 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
PFAM info
Representative CDS sequence
>Lus10027968 pacid=23181264 polypeptide=Lus10027968 locus=Lus10027968.g ID=Lus10027968.BGIv1.0 annot-version=v1.0
ATGTTCTCCACCGTACAGAATCTGTACCTGAACAATAACCGGTTCAGCGGTGAGGTCCCCGGGAGTCTTGTAGACCGGTTGATGGCGACAGGGATGGAGA
CTCTGTATTTACAGCATAACTATCTGACGGGGATCGAGATTAATCCGACGGCGGAGATTCCGGTGAGTAGCGCGCTGTGTTTGCAGTACAATTGCATGGT
GCCGCCGGTGCAGACTCCTTGCCCGTTGAAGGCCGGGCAGCAGAAGACTAGGCCGACCGATCAATGCAGAGAATGGAGAGGGTAA
AA sequence
>Lus10027968 pacid=23181264 polypeptide=Lus10027968 locus=Lus10027968.g ID=Lus10027968.BGIv1.0 annot-version=v1.0
MFSTVQNLYLNNNRFSGEVPGSLVDRLMATGMETLYLQHNYLTGIEINPTAEIPVSSALCLQYNCMVPPVQTPCPLKAGQQKTRPTDQCREWRG
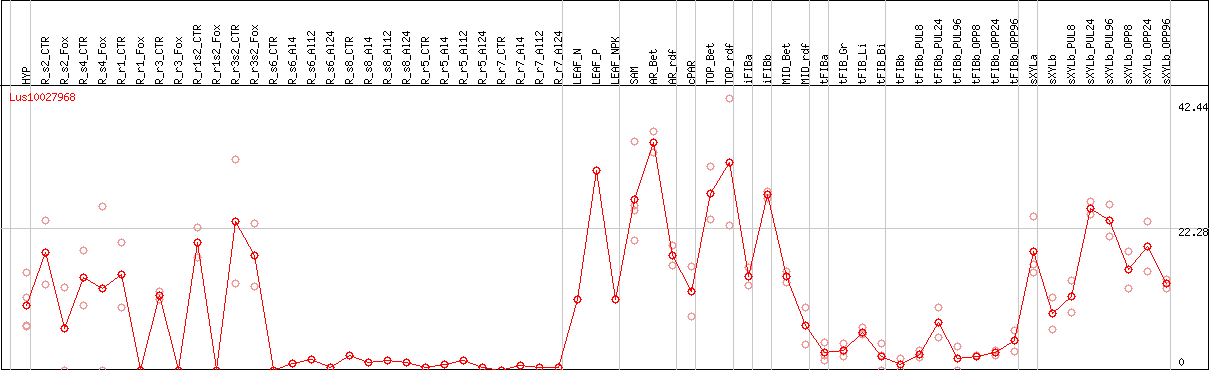
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10027968 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.