External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G60510 453 / 6e-160
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.2), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.3)
AT4G31810 447 / 1e-157
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
AT5G65940 324 / 1e-109
CHY1
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
AT2G30660 315 / 4e-106
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
AT2G30650 313 / 4e-105
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
AT1G06550 271 / 7e-89
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
AT3G24360 178 / 2e-52
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.2)
AT4G13360 178 / 3e-52
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
AT4G16800 50 / 8e-07
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
AT5G43280 45 / 3e-05
ATDCI1
"delta\(3,5\),delta\(2,4\)-dienoyl-CoA isomerase 1", "delta\(3,5\),delta\(2,4\)-dienoyl-CoA isomerase 1", delta(3,5),delta(2,4)-dienoyl-CoA isomerase 1 (.1), delta(3,5),delta(2,4)-dienoyl-CoA isomerase 1 (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042883
586 / 0
AT4G31810 394 / 4e-137
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10010685
437 / 2e-153
AT4G31810 575 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10007422
431 / 5e-151
AT4G31810 568 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10038717
310 / 8e-102
AT5G65940 565 / 0.0
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Lus10020758
256 / 2e-82
AT1G06550 602 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10007334
252 / 3e-81
AT1G06550 595 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10019396
157 / 2e-44
AT4G13360 607 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10043253
143 / 1e-38
AT4G13360 565 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Lus10010994
60 / 6e-10
AT4G16800 351 / 2e-121
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G057400
537 / 0
AT3G60510 520 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.2), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.3)
Potri.018G018800
443 / 5e-156
AT4G31810 565 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
Potri.001G156900
423 / 4e-148
AT3G60510 432 / 9e-151
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.2), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.3)
Potri.014G179000
319 / 2e-107
AT5G65940 556 / 0.0
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Potri.006G277300
280 / 5e-92
AT5G65940 397 / 1e-137
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Potri.010G170200
281 / 8e-92
AT5G65940 388 / 6e-133
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Potri.018G004200
273 / 2e-89
AT5G65940 430 / 2e-150
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Potri.018G004150
272 / 3e-89
AT5G65940 428 / 6e-150
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Potri.018G004100
269 / 8e-88
AT5G65940 393 / 6e-136
beta-hydroxyisobutyryl-CoA hydrolase 1 (.1.2.3)
Potri.002G057700
256 / 9e-83
AT1G06550 592 / 0.0
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0127
ClpP_crotonase
PF00378
ECH_1
Enoyl-CoA hydratase/isomerase
Representative CDS sequence
>Lus10028182 pacid=23166475 polypeptide=Lus10028182 locus=Lus10028182.g ID=Lus10028182.BGIv1.0 annot-version=v1.0
ATGGGCGCTCGATTGAACAGATTGTACAAATCGTGGGAAGATGATCCTAATGTTGGATTTGTGGCCATGAAGGGCAGTGGCAGGGCATTTTGCGCTGGTG
GGGATATTGTTTCTCTTCATCACTTCATAAACCAGGGGAAGTTCGATAGCTGCAAAGAATTCTTTTCTACATTGTACAGTTATATATACCTGCTAGGTAC
ATACTTGAAGCCAAACGTTGCTCTACTGAATGGCATCACTATGGGTGGTGGTGCTGGAGTTTCGATTCCTGGGATGTTTAGGGTTGCGACTGAAAGAACT
TTATTTGCTACTCCTGAAACCCTTATTGGTTTTCACCCTGATGCTGGTGCATCATTCTACCTTTCACATTTACCTGGTCATTTAGGTGAATATCTTGCTC
TTACGGGTGAGACACTGAAGGGAGCTGAAATGATTGGTTGTGGCTTAGCTACTCATTACACTCACAGTACAAAGCTTGGATTAGTTGAGAGAAAGTTGGC
TGATCTGGCCACTGATGATCCTTCTGTAATTGAAACTTCATTAGAAGAATATGGTGACATTGTCCACCCAGATAACATGAGTGTACTTCACAGGATTGAA
ACGGTTGATAAATGTTTCAGCCTGAACACAGTTGAAGAAATCATAGATGCTTTGGAAACTGAAGCTGCAGCAACGAATGATCCCTGGTGCAATTCAACAC
TGAGAAGACTCGGAGAATGCTCCCCATTGAGTTTAAAGGTTTCATTAAAATCTATACGAGATGGAAGATTTCAGACATTTGATCAGTGCTTAGTCCGTGA
ATATAGAATGTCTCTACAGGGGATCTCCCAGCCAGACTTCTGCGAGGGTGTTAGGGCACGTATGGTAGACAAGGACTTGGCTGCAAAGGTACGGTGGAAT
CCCCCAAGGTTGGAGCAGGTGACAGATGACATGGTTGATCGTTATTTCGCTCCGCTAAGTGAATCTGAACCGGATCTGGAGCTTCCGACGCAGCAGCGGG
AAGCATTTACCTCATAA
AA sequence
>Lus10028182 pacid=23166475 polypeptide=Lus10028182 locus=Lus10028182.g ID=Lus10028182.BGIv1.0 annot-version=v1.0
MGARLNRLYKSWEDDPNVGFVAMKGSGRAFCAGGDIVSLHHFINQGKFDSCKEFFSTLYSYIYLLGTYLKPNVALLNGITMGGGAGVSIPGMFRVATERT
LFATPETLIGFHPDAGASFYLSHLPGHLGEYLALTGETLKGAEMIGCGLATHYTHSTKLGLVERKLADLATDDPSVIETSLEEYGDIVHPDNMSVLHRIE
TVDKCFSLNTVEEIIDALETEAAATNDPWCNSTLRRLGECSPLSLKVSLKSIRDGRFQTFDQCLVREYRMSLQGISQPDFCEGVRARMVDKDLAAKVRWN
PPRLEQVTDDMVDRYFAPLSESEPDLELPTQQREAFTS
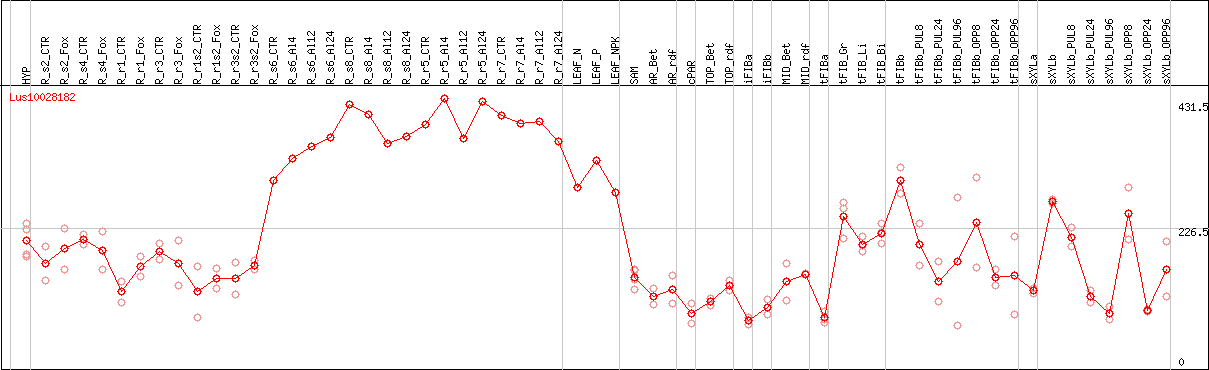
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028182 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.