External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G51060 205 / 3e-63
SRS1, STY1
STYLISH 1, SHI RELATED SEQUENCE 1, Lateral root primordium (LRP) protein-related (.1)
AT5G66350 194 / 1e-59
SHI
SHORT INTERNODES, Lateral root primordium (LRP) protein-related (.1)
AT1G75520 173 / 3e-51
SRS5
SHI-related sequence 5 (.1)
AT4G36260 167 / 4e-49
SRS2, STY2
STYLISH 2, SHI RELATED SEQUENCE 2, Lateral root primordium (LRP) protein-related (.1)
AT1G19790 166 / 2e-48
SRS7
SHI-related sequence 7 (.1.2)
AT2G18120 147 / 1e-42
SRS4
SHI-related sequence 4 (.1)
AT2G21400 132 / 2e-37
SRS3
SHI-related sequence3 (.1)
AT5G12330 127 / 4e-34
LRP1
LATERAL ROOT PRIMORDIUM 1, Lateral root primordium (LRP) protein-related (.1), Lateral root primordium (LRP) protein-related (.2), Lateral root primordium (LRP) protein-related (.3), Lateral root primordium (LRP) protein-related (.4)
AT3G54430 92 / 3e-22
SRS6
SHI-related sequence 6 (.1)
AT5G33210 88 / 2e-21
SRS8
SHI-related sequence 8 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041803
196 / 5e-63
AT3G51060 88 / 1e-21
STYLISH 1, SHI RELATED SEQUENCE 1, Lateral root primordium (LRP) protein-related (.1)
Lus10024305
178 / 7e-53
AT1G75520 260 / 8e-85
SHI-related sequence 5 (.1)
Lus10003416
171 / 2e-50
AT1G75520 258 / 6e-84
SHI-related sequence 5 (.1)
Lus10041802
134 / 3e-37
AT5G66350 150 / 4e-44
SHORT INTERNODES, Lateral root primordium (LRP) protein-related (.1)
Lus10036032
131 / 3e-36
AT5G12330 214 / 2e-68
LATERAL ROOT PRIMORDIUM 1, Lateral root primordium (LRP) protein-related (.1), Lateral root primordium (LRP) protein-related (.2), Lateral root primordium (LRP) protein-related (.3), Lateral root primordium (LRP) protein-related (.4)
Lus10009697
91 / 5e-22
AT5G66350 103 / 5e-27
SHORT INTERNODES, Lateral root primordium (LRP) protein-related (.1)
Lus10017492
87 / 4e-20
AT2G21400 120 / 1e-34
SHI-related sequence3 (.1)
Lus10028794
81 / 7e-18
AT5G66350 123 / 5e-34
SHORT INTERNODES, Lateral root primordium (LRP) protein-related (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G017500
212 / 2e-66
AT3G51060 238 / 6e-76
STYLISH 1, SHI RELATED SEQUENCE 1, Lateral root primordium (LRP) protein-related (.1)
Potri.005G118200
211 / 7e-66
AT3G51060 233 / 6e-74
STYLISH 1, SHI RELATED SEQUENCE 1, Lateral root primordium (LRP) protein-related (.1)
Potri.002G028500
177 / 4e-53
AT1G75520 261 / 1e-85
SHI-related sequence 5 (.1)
Potri.005G234200
162 / 2e-47
AT1G75520 222 / 2e-70
SHI-related sequence 5 (.1)
Potri.004G160600
156 / 3e-46
AT5G66350 170 / 4e-52
SHORT INTERNODES, Lateral root primordium (LRP) protein-related (.1)
Potri.009G121600
150 / 5e-44
AT5G66350 181 / 3e-56
SHORT INTERNODES, Lateral root primordium (LRP) protein-related (.1)
Potri.009G070800
131 / 2e-35
AT5G12330 231 / 9e-74
LATERAL ROOT PRIMORDIUM 1, Lateral root primordium (LRP) protein-related (.1), Lateral root primordium (LRP) protein-related (.2), Lateral root primordium (LRP) protein-related (.3), Lateral root primordium (LRP) protein-related (.4)
Potri.001G276200
131 / 4e-35
AT5G12330 226 / 2e-71
LATERAL ROOT PRIMORDIUM 1, Lateral root primordium (LRP) protein-related (.1), Lateral root primordium (LRP) protein-related (.2), Lateral root primordium (LRP) protein-related (.3), Lateral root primordium (LRP) protein-related (.4)
Potri.003G196100
128 / 9e-35
AT5G12330 159 / 8e-47
LATERAL ROOT PRIMORDIUM 1, Lateral root primordium (LRP) protein-related (.1), Lateral root primordium (LRP) protein-related (.2), Lateral root primordium (LRP) protein-related (.3), Lateral root primordium (LRP) protein-related (.4)
Potri.001G027700
121 / 3e-32
AT5G12330 157 / 4e-46
LATERAL ROOT PRIMORDIUM 1, Lateral root primordium (LRP) protein-related (.1), Lateral root primordium (LRP) protein-related (.2), Lateral root primordium (LRP) protein-related (.3), Lateral root primordium (LRP) protein-related (.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05142
DUF702
Domain of unknown function (DUF702)
Representative CDS sequence
>Lus10028352 pacid=23166175 polypeptide=Lus10028352 locus=Lus10028352.g ID=Lus10028352.BGIv1.0 annot-version=v1.0
ATGGCTGGTCTCTTCTCTTTAGGCGGTGGTGGTGGAGGAGGAGGAGGGAGGGGTAGTAGTGGAAACACAAATAATAATAATACAGAGATCACGCCGGAGA
GCAATAACAACAGACAGCAGCAGCAGCAGGCGGAATTGATGCAGCGGCACCAAGATCTCTACTCTTCCGCTGCAGGGTTAGGAGTGGGACCCACTCATCA
TCACTTGTACCTATCTGCTGATCATCACCACCACCACCCTTCTTCCTCCAGATCCGCCGCCGCGTTAATGATGATGCGTCCCGGCGGAGGAGAAGGAGGA
AGGGGCTCCAGCAGTAGTATTAGCAACGTTGGTGGTGGAATTAGTTGCCAAGACTGTGGGAACCAAGCGAAAAAGGACTGCAGCCATAGTAGATGCAGGA
CTTGCTGCAAGAGCCGCGGCTACAATTGCCAAACCCATGTCAAAAGCACTTGGGTTCCTGCCGCCAAACGCCGTGAGCGCCAGCAGCAGCAGCACCCCGT
TACCTCCGTCCAACAGCCACATCAGCAGCAGCATCAAGTTGTTCCCAAGCGGCCTAGAGATACAACATCTTCAACTCTCGTCTCCCATCCTCCTGGGTTA
GAGGTGGCGGTAGGTAATTTCCCGGCAGAAGTGAGCTCGGAGGCGCTGCTCCGGTGCGTGAGGATGAGCGGAGTGGACGAGAGCGATGCACAGCAGTACG
CGTACCAGACGGCGGTGAGCATTGGAGGGCACACGTTCAAAGGGATACTGTACGATCAAGGTCCAGCAGGAGAGAGCTCTAGTTACATGGCGGCAACTAC
CGACACTTCATCAGCCAGTGGTGTCCAACCGCTCAATCTCATTACCACTGCCGCCTCGGCCTCAGCGTCTGCGTCCTCCAACATCGGCGGCGGTGGAGTG
ACGTCTAATGCCGCCGGAGCTAGCTACTTGGATCCGTCTTCGCTGTACACAGCCGTACCGCTAAGTAACTACATGGCTGGTACGCAATTCTTCCCGAATA
ACCCGAGATGA
AA sequence
>Lus10028352 pacid=23166175 polypeptide=Lus10028352 locus=Lus10028352.g ID=Lus10028352.BGIv1.0 annot-version=v1.0
MAGLFSLGGGGGGGGGRGSSGNTNNNNTEITPESNNNRQQQQQAELMQRHQDLYSSAAGLGVGPTHHHLYLSADHHHHHPSSSRSAAALMMMRPGGGEGG
RGSSSSISNVGGGISCQDCGNQAKKDCSHSRCRTCCKSRGYNCQTHVKSTWVPAAKRRERQQQQHPVTSVQQPHQQQHQVVPKRPRDTTSSTLVSHPPGL
EVAVGNFPAEVSSEALLRCVRMSGVDESDAQQYAYQTAVSIGGHTFKGILYDQGPAGESSSYMAATTDTSSASGVQPLNLITTAASASASASSNIGGGGV
TSNAAGASYLDPSSLYTAVPLSNYMAGTQFFPNNPR
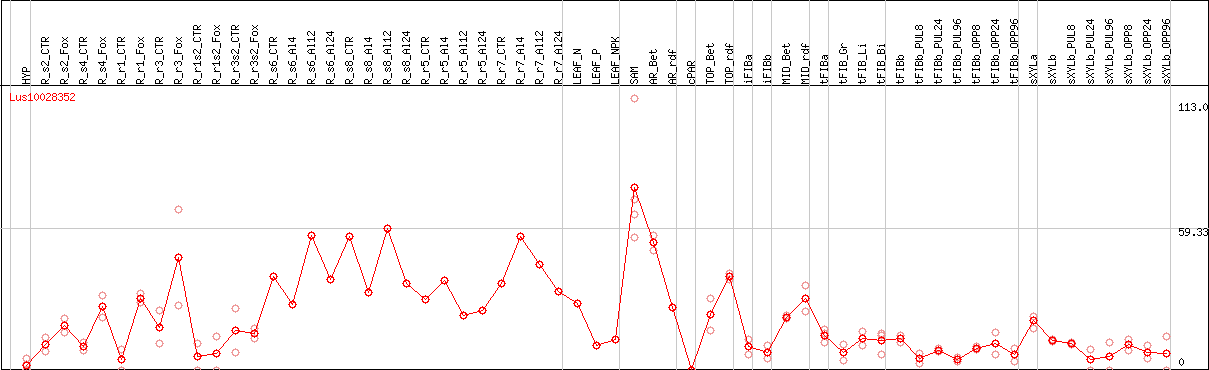
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028352 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.