External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35783 59 / 1e-13
RTFL6, DVL17
DEVIL 17, ROTUNDIFOLIA like 6 (.1)
AT2G29125 47 / 2e-08
RTFL2, DVL13
DEVIL 13, ROTUNDIFOLIA like 2 (.1)
AT1G68825 45 / 2e-08
DVL5, RTFL15
DEVIL 5, ROTUNDIFOLIA like 15 (.1.2)
AT3G46613 46 / 9e-08
RTFL4, DVL15
DEVIL 15, ROTUNDIFOLIA like 4 (.1)
AT5G59510 45 / 2e-07
RTFL5, DVL18
DEVIL 18, ROTUNDIFOLIA like 5 (.1)
AT3G55515 44 / 2e-07
DVL8, RTFL7
DEVIL 8, ROTUNDIFOLIA like 7 (.1)
AT1G13245 42 / 3e-07
RTFL17, DVL4
DEVIL 4, ROTUNDIFOLIA like 17 (.1)
AT4G13395 42 / 5e-07
DVL10, RTFL12
DEVIL 10, ROTUNDIFOLIA like 12 (.1)
AT1G67265 42 / 6e-07
DVL3, RTFL21
DEVIL 3, ROTUNDIFOLIA like 21 (.1)
AT2G39705 42 / 8e-07
RTFL8, DVL11
DEVIL 11, ROTUNDIFOLIA like 8 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G226250
49 / 7e-10
AT2G36985 67 / 3e-17
ROTUNDIFOLIA4, DEVIL 16, DVL family protein (.1)
Potri.008G035900
49 / 8e-10
AT2G36985 66 / 1e-16
ROTUNDIFOLIA4, DEVIL 16, DVL family protein (.1)
Potri.001G242800
49 / 1e-09
AT2G29125 71 / 4e-17
DEVIL 13, ROTUNDIFOLIA like 2 (.1)
Potri.009G034300
47 / 2e-08
AT2G29125 64 / 9e-15
DEVIL 13, ROTUNDIFOLIA like 2 (.1)
Potri.010G201700
45 / 6e-08
AT2G39705 72 / 2e-18
DEVIL 11, ROTUNDIFOLIA like 8 (.1)
Potri.008G057800
45 / 6e-08
AT2G39705 84 / 8e-23
DEVIL 11, ROTUNDIFOLIA like 8 (.1)
Potri.008G116700
44 / 1e-07
AT1G13245 76 / 6e-21
DEVIL 4, ROTUNDIFOLIA like 17 (.1)
Potri.010G129600
43 / 1e-07
AT1G13245 75 / 1e-20
DEVIL 4, ROTUNDIFOLIA like 17 (.1)
Potri.004G102950
42 / 6e-07
AT1G67265 64 / 4e-16
DEVIL 3, ROTUNDIFOLIA like 21 (.1)
Potri.003G073900
42 / 1e-06
AT1G53708 81 / 9e-22
ROTUNDIFOLIA like 9 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF08137
DVL
DVL family
Representative CDS sequence
>Lus10028393 pacid=23166149 polypeptide=Lus10028393 locus=Lus10028393.g ID=Lus10028393.BGIv1.0 annot-version=v1.0
ATGTTGGTGGGTAGATGTCTGACTCGCCAAGGGAGCCAAAGGGTCGTCGTCGGCGAAGTTGACGGTGGCTCCGGCGGCGTTCCTCATCAACAACAACCTG
CAGGTTGCATGGGGACGGTGAGGGAGCAGCGTTCAAGGTTTTATGTTGCGAGGAGATGCGTCGTTATGCTTCTTTGTTGGCACAAATACGGCAAGTCTTA
G
AA sequence
>Lus10028393 pacid=23166149 polypeptide=Lus10028393 locus=Lus10028393.g ID=Lus10028393.BGIv1.0 annot-version=v1.0
MLVGRCLTRQGSQRVVVGEVDGGSGGVPHQQQPAGCMGTVREQRSRFYVARRCVVMLLCWHKYGKS
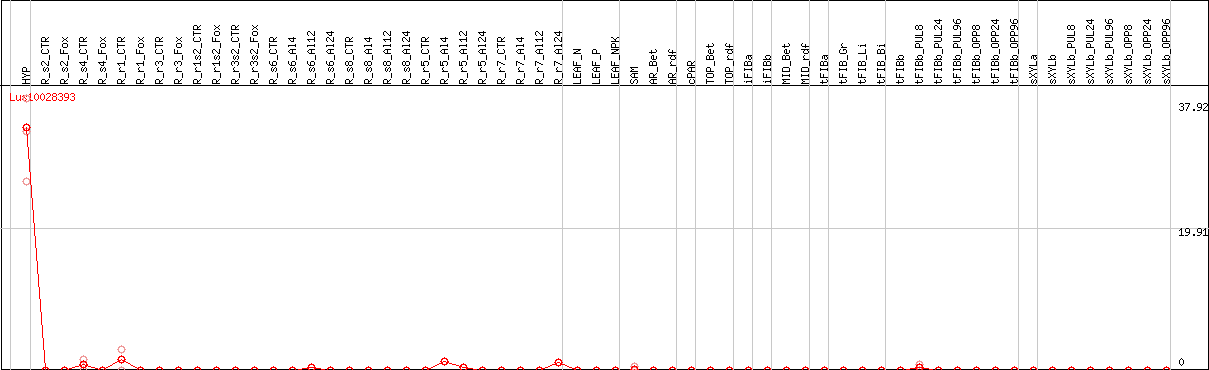
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028393 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.