External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G36020 120 / 2e-32
CSDP1
cold shock domain protein 1 (.1)
AT2G17870 117 / 3e-31
ATCSP3
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
AT3G42860 70 / 1e-13
zinc knuckle (CCHC-type) family protein (.1)
AT2G21060 65 / 2e-12
ATCSP4, ATGRP2B
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
AT1G75560 56 / 4e-09
zinc knuckle (CCHC-type) family protein (.1), zinc knuckle (CCHC-type) family protein (.2)
AT4G38680 49 / 4e-07
ATCSP2, CSDP2, GRP2
COLD SHOCK DOMAIN PROTEIN 2, ARABIDOPSIS THALIANA COLD SHOCK PROTEIN 2, glycine rich protein 2 (.1)
AT5G52380 47 / 3e-06
VASCULAR-RELATED NAC-DOMAIN 6 (.1)
AT3G43590 47 / 1e-05
zinc knuckle (CCHC-type) family protein (.1)
AT3G53500 45 / 2e-05
RSZ32, RS2Z32, At-RS2Z
arginine/serine-rich zinc knuckle-containing protein 32, RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.1), RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.2)
AT2G37340 44 / 6e-05
RSZ33, ATRSZ33, RS2Z33, AT-RS2Z33
arginine/serine-rich zinc knuckle-containing protein 33 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041902
310 / 2e-107
AT4G36020 159 / 2e-47
cold shock domain protein 1 (.1)
Lus10028073
76 / 2e-16
AT4G36020 115 / 3e-31
cold shock domain protein 1 (.1)
Lus10025058
70 / 2e-15
AT2G21060 112 / 6e-33
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Lus10005066
61 / 2e-11
AT2G17870 122 / 1e-35
ARABIDOPSIS COLD SHOCK DOMAIN PROTEIN 3, cold shock domain protein 3 (.1)
Lus10027839
61 / 2e-11
AT2G21060 110 / 6e-31
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Lus10012906
60 / 2e-10
AT3G42860 184 / 1e-54
zinc knuckle (CCHC-type) family protein (.1)
Lus10025623
57 / 4e-10
ND 81 / 8e-20
Lus10034487
57 / 5e-10
AT2G21060 110 / 7e-31
COLD SHOCK DOMAIN PROTEIN 4, glycine-rich protein 2B (.1)
Lus10030560
57 / 3e-09
AT3G42860 258 / 1e-82
zinc knuckle (CCHC-type) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0021
OB
PF00313
CSD
'Cold-shock' DNA-binding domain
CL0511
Retroviral_zf
PF00098
zf-CCHC
Zinc knuckle
Representative CDS sequence
>Lus10028449 pacid=23166123 polypeptide=Lus10028449 locus=Lus10028449.g ID=Lus10028449.BGIv1.0 annot-version=v1.0
ATGGCGGAAGATCAGTCAGTTAGATCTTCATCCACAGTGAGGACCACCGGCAAGGTCGTCAAGTTCAGCGACAAGAAGGGTTTCGGCTTCATCAAGCCCA
ACGATGGCAGCGACGACTTGTTCGTCCATCACACCGCGATCAAGTCCGACGGCGGGTACCGCACCCTCACTGAGGAAGATGTAGTCGAGTTCGAAGTAGC
CGAGACTGACGGCAAGTGCCAGGCTATCAACGTCACTGCTCCCGGCGGAGGTGCGATTCAGGCGGGGAAACGAGGCGGAGGGGACGGTTACGGCAGGCGC
GGTGCGGGATCCGGAGCGCCGTGGAACAGGCGGAGCAACGGCGGCGGGTGTTACACCTGCGGAAACCCTAACCACATCGCGAGGGAGTGCAACAGTCGCA
GCGAGAGCGTAGGCAGCTGCTTCAAGTGTGGGACCGCTGGGCATTTTGCTCGGGACTGTCCGCAGGCCGTTAACGGCCGTGGTGGCGGAGGAGTTGGCGG
AGGAGCTGCTGGTGTGGCTTGCTTTAGCTGCAGTGGATTCGGACACTTGGCTAGGGATTGTCCAGGCGGCAGCGGACCTTGCTACAACTGTGGGGGTCAT
GGACACTTCGCTAGAGATTGTACCAGCGCTAGGAGCGTGGGCGGTGGCGGTGGTGGCGGCAACAGGGTTGGGTCTAACAGTGCTTGCTTCAATTGCGGGA
AGCTAGGACACTTTGCAAGGGATTGTACCAACCCTGACTTTTGA
AA sequence
>Lus10028449 pacid=23166123 polypeptide=Lus10028449 locus=Lus10028449.g ID=Lus10028449.BGIv1.0 annot-version=v1.0
MAEDQSVRSSSTVRTTGKVVKFSDKKGFGFIKPNDGSDDLFVHHTAIKSDGGYRTLTEEDVVEFEVAETDGKCQAINVTAPGGGAIQAGKRGGGDGYGRR
GAGSGAPWNRRSNGGGCYTCGNPNHIARECNSRSESVGSCFKCGTAGHFARDCPQAVNGRGGGGVGGGAAGVACFSCSGFGHLARDCPGGSGPCYNCGGH
GHFARDCTSARSVGGGGGGGNRVGSNSACFNCGKLGHFARDCTNPDF
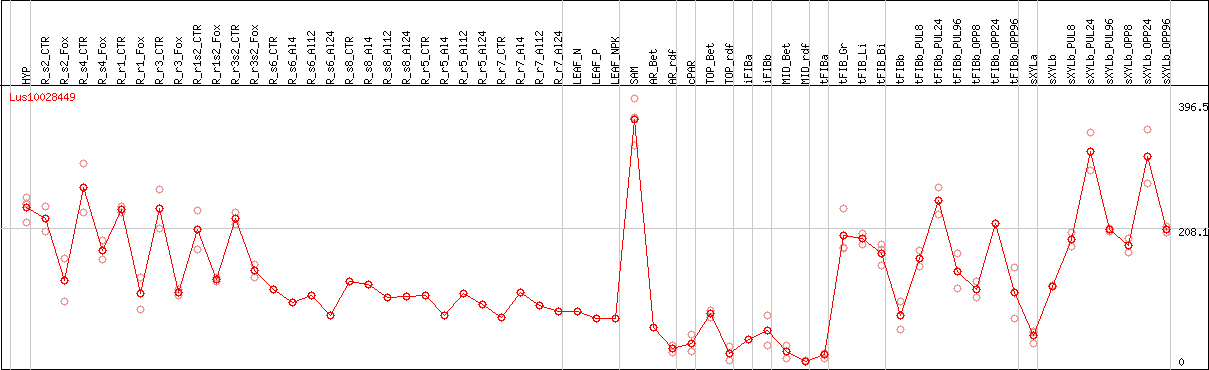
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10028449 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.